The concept of the “base pairing rule” has long been the cornerstone of molecular biology, explaining how life encodes, replicates, and transmits information. However, in the modern era of the Fourth Industrial Revolution, this biological principle has transcended the laboratory. Today, the base pairing rule serves as a critical blueprint for the most advanced frontiers of technology, including DNA-based data storage, bioinformatics, AI-driven drug discovery, and molecular computing.
By understanding the “if-then” logic inherent in adenine (A) pairing with thymine (T) and cytosine (C) pairing with guanine (G), technologists are developing systems that can store the world’s data for centuries or compute complex problems at a molecular scale. This article explores the base pairing rule through the lens of technology, examining how this ancient biological algorithm is reshaping the future of digital infrastructure.
Decoding the Biological Framework for Tech Application
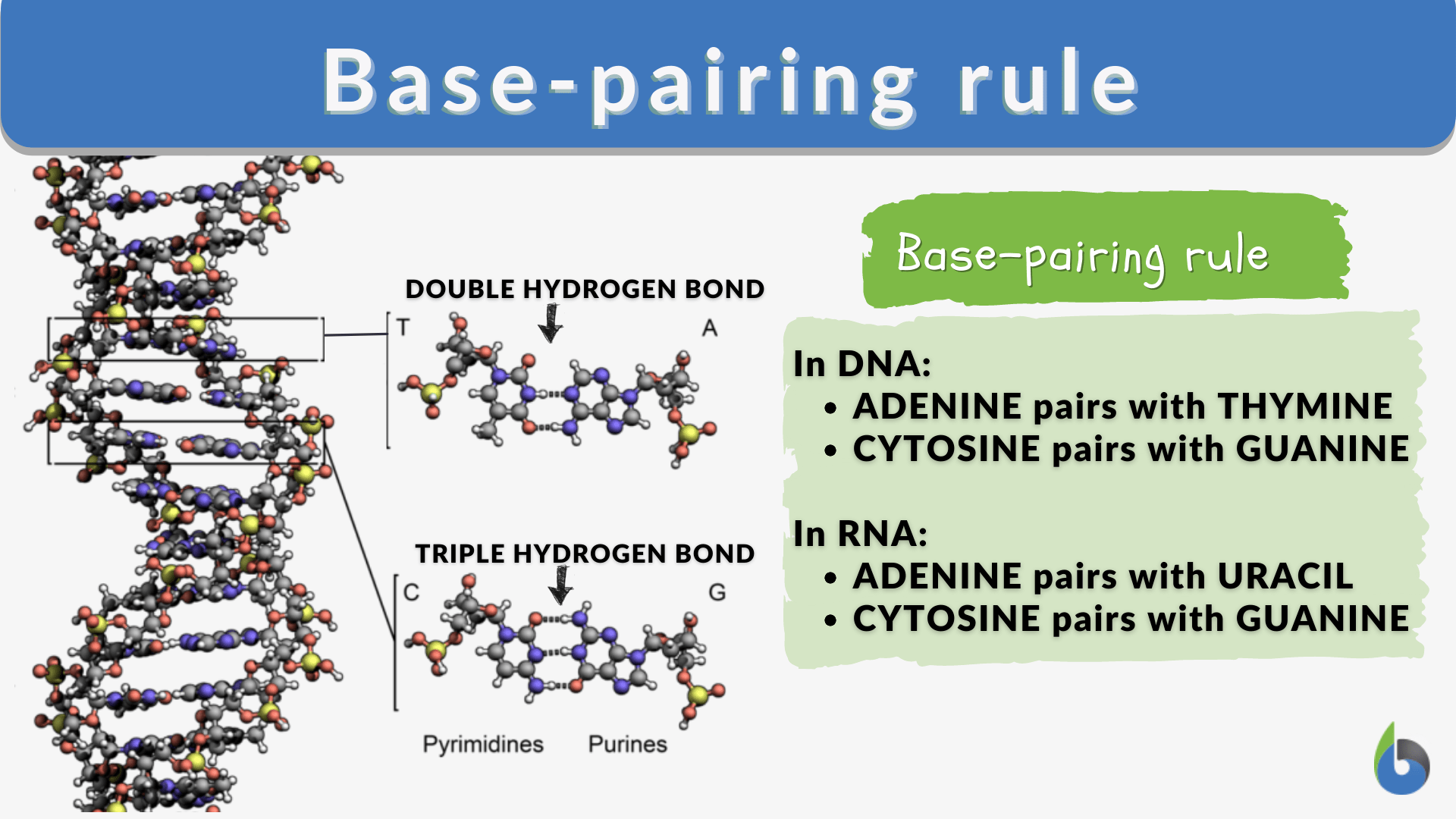
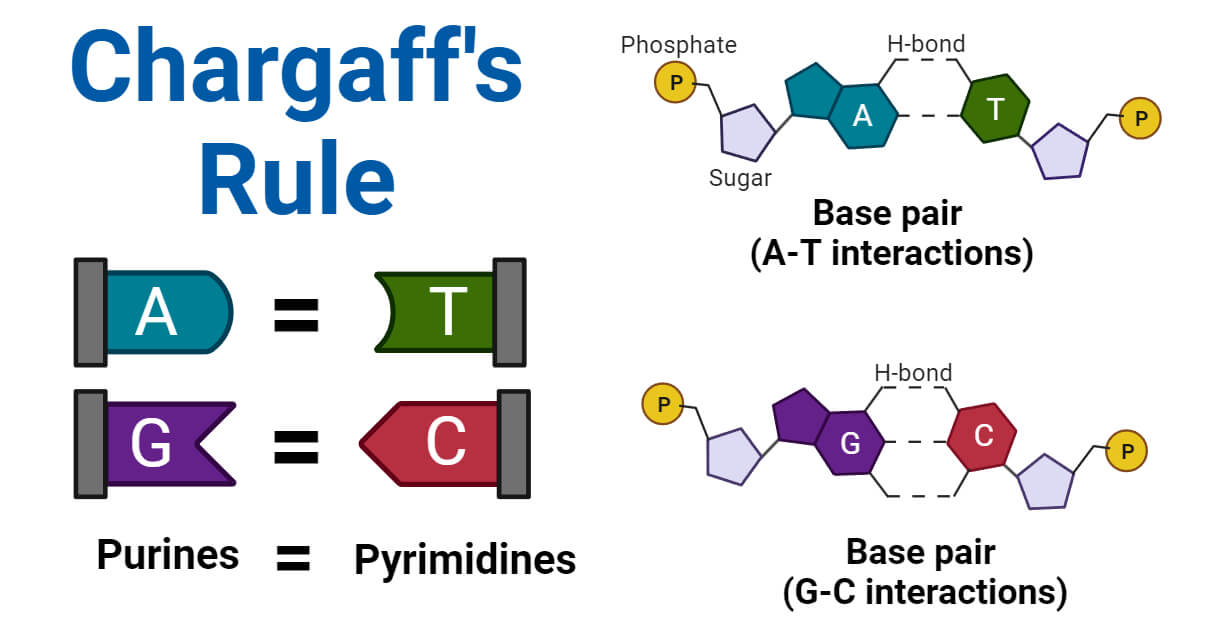
To understand how the base pairing rule influences technology, one must first grasp its inherent logic. In the double-helix structure of DNA, the nitrogenous bases do not bond at random. They follow a strict, complementary protocol: Adenine always pairs with Thymine (forming two hydrogen bonds), and Cytosine always pairs with Guanine (forming three hydrogen bonds).
The Fundamental Logic of Complementarity
In the world of computer science, we operate on binary code—zeros and ones. In the biological world, the base pairing rule provides a quaternary (four-base) system. This biological complementarity acts as a natural error-correction mechanism. Because one strand of DNA dictates the exact sequence of its partner, the system possesses built-in redundancy. For technologists, this is the ultimate template for data integrity. If a sequence is damaged or corrupted, the “complementary rule” allows for the perfect reconstruction of the lost information, a concept that is currently being integrated into high-end digital security and data recovery protocols.
Precision and Error Correction in Molecular Coding
Software developers often struggle with “bit rot” or data degradation in traditional magnetic and optical storage. The base pairing rule offers a solution through its chemical specificity. By leveraging the hydrogen bonding affinity between A-T and C-G, bio-engineers are designing “smart” molecular probes and sensors. These tools use the base pairing rule to identify specific digital signatures within a sea of biological data, ensuring that “noise” does not interfere with the signal—a principle directly mirrored in digital signal processing and error-correcting codes used in 5G and satellite communications.
DNA Data Storage: The Future of High-Density Archiving
One of the most disruptive applications of the base pairing rule in tech is DNA Data Storage. As the world produces an exponential amount of data, traditional silicon-based storage is reaching its physical limits. DNA, governed by the base pairing rule, offers a density millions of times greater than any flash drive or hard disk.
Translating Binary to Quaternary Systems
To store a movie, a document, or a codebase in DNA, engineers must first translate the digital binary (0s and 1s) into the biological quaternary code (A, T, C, G). The base pairing rule ensures that when these synthetic strands are synthesized, they can be reliably “read” by sequencing machines. For example, a “00” might be represented by Adenine, while “11” is represented by Guanine. Because we know that A will only ever pair with T during the replication process (PCR), we can amplify this data billions of times without losing the original digital sequence. This enables the archiving of petabytes of data in a space no larger than a sugar cube.
Solving the Longevity Crisis of Traditional Hardware
Technology trends are currently shifting toward “cold storage” solutions—data that needs to be kept safe for hundreds of years without power. Magnetic tapes degrade in decades; DNA, when kept in a cool, dark environment, can last for millennia. The stability of the base pairing rule is what makes this possible. Unlike a hard drive that might suffer from mechanical failure or a cloud server requiring constant electricity, a DNA-based “hard drive” relies on the chemical stability of the A-T and C-G bonds. Tech giants like Microsoft and startups like Twist Bioscience are already utilizing these rules to create the next generation of archival technology.
Bioinformatics and AI: Algorithms Mimicking Nature
The intersection of artificial intelligence and biology is perhaps where the base pairing rule is most influential. Bioinformatics—the use of software tools to understand biological data—relies entirely on the predictability of base pairing to sequence genomes and predict molecular behavior.
Pattern Recognition in Genomic Sequencing
Modern sequencing technologies, such as Next-Generation Sequencing (NGS), use the base pairing rule as their operational “software.” When a sequencer identifies a strand of DNA, it uses fluorescently labeled nucleotides that bind to their complementary partners. AI algorithms then analyze these flashes of light to reconstruct the genome. Without the mathematical certainty of the base pairing rule, the software would be unable to align the millions of short fragments of data into a coherent map. This is essentially a massive data-matching exercise, similar to how search engines index the internet, but at a molecular level.
Machine Learning for Protein Folding and Synthesis
AI tools like Google DeepMind’s AlphaFold have revolutionized how we understand life by predicting how proteins fold. This process begins with the DNA sequence, where the base pairing rule determines the messenger RNA, which in turn determines the protein. By feeding the rules of base pairing into deep learning models, tech companies can now “program” new proteins or enzymes for industrial use, such as plastic-eating bacteria or high-efficiency biofuels. The AI doesn’t just see letters; it sees the underlying chemical logic of the pairings, allowing it to simulate biological outcomes with 99% accuracy.
Nanotechnology and DNA Computing
While we usually think of DNA as a storage medium, the tech industry is increasingly viewing it as a processor. This field, known as DNA computing or molecular programming, uses the base pairing rule to perform logical operations.
Logic Gates Built from Nucleotides
In a traditional computer, logic gates (AND, OR, NOT) are made of transistors. In a DNA computer, these gates are made of DNA strands. By utilizing the base pairing rule, researchers can create “input” strands that only bind to “output” strands if a specific sequence is present. This is a form of massive parallel processing. Because trillions of DNA strands can pair up simultaneously in a single test tube, a DNA computer could theoretically solve “unbreakable” cryptographic problems that would take a silicon supercomputer years to process.
Structural DNA Origami in Micro-Engineering
The base pairing rule also allows for “DNA Origami,” a technique in nanotechnology where DNA is folded into specific shapes to create microscopic structures. Because A always seeks T and C always seeks G, engineers can design sequences that self-assemble into nanobots, drug-delivery vehicles, or even microscopic circuit boards. This is “bottom-up” manufacturing, where the software (the DNA sequence) creates its own hardware (the physical structure) through the simple, reliable rule of base pairing.
The Security Implications of Genetic Information Systems
As we integrate the base pairing rule into our technological ecosystem, digital security becomes a paramount concern. If we are storing sensitive corporate or national data in DNA, we must develop new methods to protect it.
Cryptography Using Biological Keys
The complexity of the base pairing rule provides a new layer for cryptography. “Bio-cryptography” uses DNA sequences as encryption keys. Because of the vast permutations possible within a four-base system, a key based on a specific DNA sequence is nearly impossible to brute-force. To “unlock” the data, one would need the exact complementary strand, governed by the base pairing rule, to bind with and reveal the encrypted message. This merges the physical world of chemistry with the digital world of cybersecurity.

Ethical Tech: Protecting the Ultimate Personal Data
The move toward DNA-based technology also necessitates a discussion on digital ethics and security tools. As our “code” (our DNA) becomes readable and writeable like software, the tech industry must develop robust firewalls against “bio-hacking.” The base pairing rule is the language of this new frontier. Understanding it is not just for biologists; it is for the security analysts, software engineers, and privacy advocates who will navigate a future where the line between the biological and the digital is permanently blurred.
In conclusion, the base pairing rule is no longer confined to the realm of biology. It is a fundamental logic gate for the next generation of technology. From the way we archive our digital legacy to the way we build the computers of the future, the simple, elegant pairing of A-T and C-G is proving to be the most resilient and efficient “tech” ever discovered. By mastering this biological rule, we are unlocking a future of limitless storage, unprecedented computing power, and a new era of bio-digital synergy.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.