In the burgeoning field of bioinformatics and computational biology, the bridge between hardware and wetware is narrowing. To the software engineer or data scientist entering the biotechnology space, the four nitrogenous bases—Adenine, Guanine, Cytosine, and Thymine—are more than just biological components; they are the fundamental bits of the world’s most complex operating system. To understand how to manipulate, sequence, and store data within DNA, one must first master the hardware architecture of these molecules. This begins with the fundamental distinction between the two families of nitrogenous bases: purines and pyrimidines.

Just as a computer architect must understand the physical properties of silicon and transistors to optimize a processor, a bioinformatics specialist must understand the molecular geometry of these bases. The differences between purines and pyrimidines dictate the structural stability of the double helix, the error rates in sequencing algorithms, and the future potential of DNA-based data storage.
The Architecture of Genetic Data: Purines vs. Pyrimidines
At the most basic level, the difference between a purine and a pyrimidine is one of structural complexity and atomic “footprint.” In the context of technology and data modeling, this is equivalent to understanding the difference between a high-density and a low-density storage format.
Chemical Structures and Encoding (Double Rings vs. Single Rings)
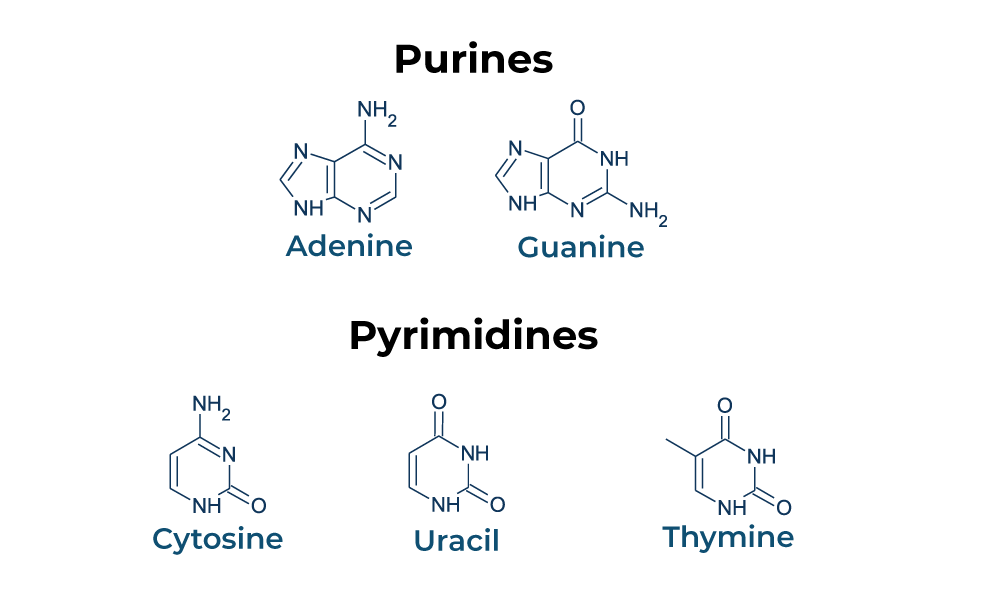
Purines are the “heavyweights” of the genetic code. They consist of a double-ring structure—a six-membered pyrimidine ring fused to a five-membered imidazole ring. In the DNA and RNA “alphabet,” the purines are Adenine (A) and Guanine (G). Because of their double-ring structure, purines are physically larger than their counterparts.
Pyrimidines, conversely, are smaller. They consist of a single six-membered nitrogenous ring. In DNA, the pyrimidines are Cytosine (C) and Thymine (T); in RNA, Uracil (U) replaces Thymine. From a tech perspective, this size disparity is critical. The logic of DNA is built on the pairing of a large purine with a small pyrimidine. If the “system” attempted to pair two purines together, the resulting “data track” (the double helix) would bulge; if it paired two pyrimidines, the track would narrow. This consistent width is what allows DNA polymerase—the biological “read/write head”—to move along the strand with high fidelity.
Base Pairing Logic: The Binary System of Biology
The “code” of life operates on a logic of complementarity. In a standard computational binary system, we use 0 and 1. In the biological system, the interaction between purines and pyrimidines creates a quaternary system. However, the pairing rules are rigid: Adenine (purine) always pairs with Thymine (pyrimidine), and Guanine (purine) always pairs with Cytosine (pyrimidine).
This pairing is governed by hydrogen bonding. A-T pairs are held together by two hydrogen bonds, while G-C pairs are held by three. For developers working on sequencing software, this is a vital distinction. G-C rich regions of a genome are “stickier” and require more energy (higher temperatures) to unzip during the Polymerase Chain Reaction (PCR) process. This physical “hardware” reality must be accounted for in the algorithms used to simulate DNA melting points and primer design.
Bioinformatics and Computational Modeling: Leveraging Nucleotide Geometry
Modern tech thrives on the ability to turn physical phenomena into predictable data models. In bioinformatics, the structural differences between purines and pyrimidines are used to refine everything from alignment algorithms to the detection of genetic mutations.
Predictive Analysis in DNA Sequencing
When we use Next-Generation Sequencing (NGS) technologies, such as those developed by Illumina or Oxford Nanopore, we are essentially digitizing the chemical properties of purines and pyrimidines. Nanopore sequencing, for instance, works by passing a single strand of DNA through a microscopic pore and measuring the disruption in electrical current.
Because purines (A and G) are larger and have different electronic signatures than pyrimidines (C and T), each base causes a unique “wiggle” in the data stream. Tech professionals in this space develop signal-processing algorithms that can distinguish these minute electronic variations in real-time. Understanding the specific molecular weight and ring structure of a purine versus a pyrimidine is the baseline for training the neural networks that perform base-calling—the process of converting electrical signals into text-based genetic sequences.
Structural Stability and Algorithm Efficiency
The difference in hydrogen bonding between purine-pyrimidine pairs (2 bonds for A-T vs. 3 bonds for G-C) introduces a variable known as “GC-bias.” This is a significant challenge in computational biology. When sequencing a genome, areas high in the pyrimidine Cytosine and the purine Guanine are often underrepresented in the final data because they are harder to “read” physically.

Software engineers must write “bias-correction” algorithms to normalize this data. If the software does not account for the structural “grip” of G-C pairs, the resulting genomic map will have “dark regions” or gaps. By modeling the thermodynamic stability of these nitrogenous bases, tech platforms can more accurately reconstruct a person’s genome from millions of short, digital fragments.
The Role of AI and Machine Learning in Genetic Engineering
Artificial Intelligence (AI) has revolutionized how we interpret the relationship between purines and pyrimidines. We are no longer just reading the code; we are using machine learning to predict how changes in these bases will affect the “output” of the organism.
Deep Learning for Mutational Analysis
A mutation often involves the substitution of one base for another. These are categorized into “transitions” and “transversions.” A transition is a swap within the same family (e.g., a purine for a purine, A ↔ G), while a transversion is a swap between families (e.g., a purine for a pyrimidine, A ↔ C).
Machine learning models, such as DeepMind’s AlphaFold or various SNP (Single Nucleotide Polymorphism) predictors, use these categories to calculate the likelihood of a disease. Transversions are often more “disruptive” to the software of the cell because they cause a more significant change in the physical shape of the DNA. AI tools are trained to identify these structural anomalies, allowing researchers to pinpoint potential cancerous mutations in a sea of gigabytes of genetic data.
CRISPR and Precision Tech Tools
CRISPR-Cas9 is essentially a “search and replace” tool for DNA. To function, it uses a guide RNA (gRNA) to find a specific sequence of purines and pyrimidines. The tech behind CRISPR involves calculating the “binding energy” between the guide RNA and the target DNA.
Since RNA uses the pyrimidine Uracil instead of Thymine, the computational modeling of these interactions is slightly different. Tech companies in the gene-editing space develop software that predicts “off-target effects”—essentially bugs in the code where the CRISPR tool might accidentally edit the wrong sequence. These predictions are based entirely on the mathematical probability of purine-pyrimidine hydrogen bonding patterns.
Future Trends: From DNA Data Storage to Quantum Biology
As we look toward the future of technology, the distinction between purine and pyrimidine moves from the realm of biology into the realm of hardware engineering and cybersecurity.
The Rise of Molecular Computing
Silicon-based chips are reaching their physical limits. DNA data storage is the next frontier, offering a density millions of times greater than current flash drives. In this tech niche, engineers encode binary data (00, 01, 10, 11) into the four bases (A, C, G, T).
The difference between purines and pyrimidines is used here to ensure data integrity. Designers of DNA “hard drives” often use the purine/pyrimidine distinction to create error-correcting codes. For example, a “parity bit” in DNA might be represented by ensuring a specific ratio of purines to pyrimidines in a sequence. If a “read” returns too many purines, the software knows the data has been corrupted by a chemical mutation and can trigger an auto-repair protocol.
Security in Biological Data
As genetic data becomes a primary form of identification, the “digital security” of our purines and pyrimidines is paramount. We are seeing the emergence of “Bio-IT” security, where cryptographic keys are stored within synthetic DNA strands.
The structural uniqueness of the purine-pyrimidine interface makes it an ideal medium for “Physical Unclonable Functions” (PUFs). Because the exact molecular sequence of a synthetic strand is nearly impossible to guess but easy to “verify” with a sequencer, the specific arrangement of these nitrogenous bases is becoming the ultimate password.
In conclusion, the differences between purines and pyrimidines are not merely academic observations for biologists. They are the physical constraints and logical parameters of the next generation of technology. Whether it is through the development of AI-driven drug discovery, the perfection of DNA sequencing algorithms, or the engineering of biological data storage, the tech industry is increasingly reliant on understanding the elegant, double-ringed and single-ringed structures that encode the very essence of life. For the modern tech professional, the genome is the ultimate codebase, and the purines and pyrimidines are its most fundamental characters.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.