The landscape of modern technology is no longer confined to silicon chips and fiber-optic cables. We have entered an era of “biological programming,” where the code of life—DNA—is being edited with the same precision developers use to debug software. At the heart of this revolution is a small but incredibly powerful molecule known as sgRNA (single guide RNA).
As the primary navigational component of the CRISPR-Cas9 system, sgRNA has transformed genetic engineering from a hit-or-miss endeavor into a programmable high-tech discipline. To understand what sgRNA is, one must view it not just as a biological entity, but as a sophisticated targeting system that allows scientists to interface with the genome.
Understanding the Fundamentals: The Architecture of sgRNA
In nature, the CRISPR-Cas9 system is an adaptive immune response found in bacteria. It originally utilized two distinct RNA molecules to identify and neutralize viral threats. However, the breakthrough that led to the Nobel Prize in Chemistry involved the technical merging of these two molecules into a single, streamlined entity: the single guide RNA (sgRNA).
The Fusion of crRNA and tracrRNA
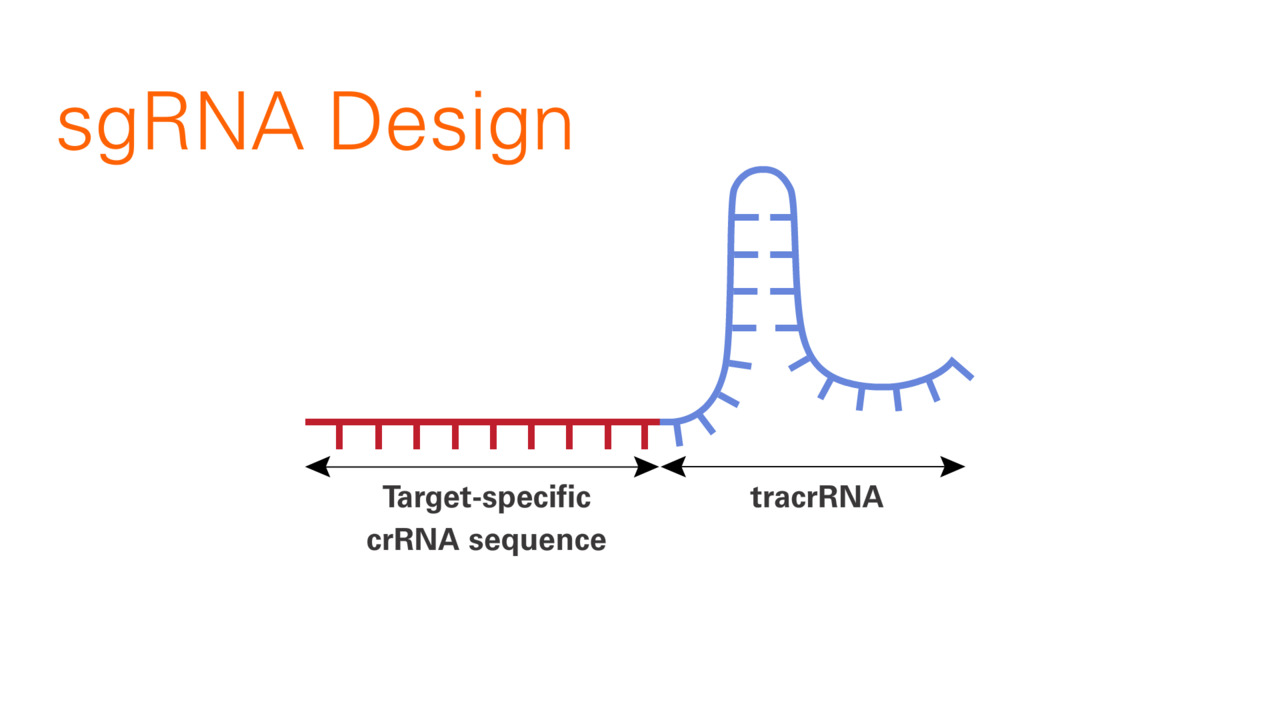
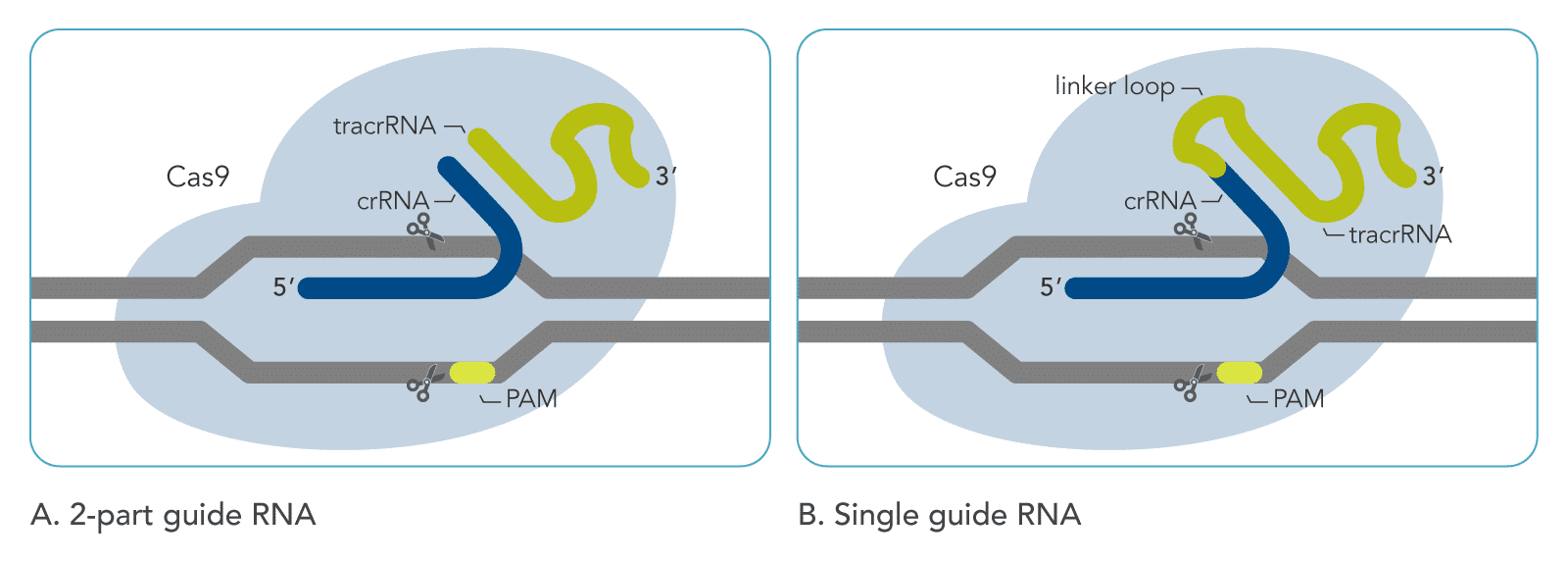
The sgRNA is a synthetic construct that combines CRISPR RNA (crRNA) and trans-activating CRISPR RNA (tracrRNA). The crRNA contains the “spacer” sequence—a 20-nucleotide string that is custom-designed to match a specific target DNA sequence. The tracrRNA acts as a structural scaffold, providing a handle for the Cas9 protein to bind to. By fusing these two into a single guide, bio-engineers created a “plug-and-play” system that drastically simplified the process of gene editing.
The Role of the Scaffold Sequence
While the 20-nucleotide targeting sequence gets most of the attention, the scaffold portion of the sgRNA is a masterpiece of molecular engineering. This constant region forms a complex three-dimensional structure that fits perfectly into the “cleft” of the Cas9 enzyme. Without this structural stability, the enzyme would be unable to maintain its shape or exert its catalytic function. In the world of tech, if the crRNA is the software code, the scaffold is the hardware interface that allows the code to execute on the processor (Cas9).
The Mechanics of Precision: How sgRNA Directs CRISPR-Cas9
The primary function of sgRNA is to act as a molecular GPS. In the vast “database” of a genome—which in humans consists of over three billion base pairs—finding a single specific gene to edit is an immense computational and physical challenge. The sgRNA solves this by leveraging the principles of Watson-Crick base pairing to achieve unparalleled specificity.
Target Recognition and Base Pairing
When a Cas9 protein is loaded with an sgRNA, it scans the genome, unzipping the DNA double helix in short bursts. The sgRNA’s targeting sequence attempts to pair with the exposed DNA. If the sequence matches, the sgRNA binds to the DNA, forming an RNA-DNA heteroduplex. This binding triggers a conformational change in the Cas9 enzyme, signaling its “molecular scissors” to activate and create a double-strand break in the DNA. This process is highly efficient, allowing for precise deletions, insertions, or modifications at the exact coordinates specified by the guide.
Navigating the PAM Sequence
Precision is further reinforced by a technical requirement known as the Protospacer Adjacent Motif (PAM). For the sgRNA to successfully bind and Cas9 to cut, a specific short DNA sequence (usually NGG for the most common Cas9) must be immediately adjacent to the target site. The PAM acts as an authentication key; it prevents the CRISPR system from attacking the bacteria’s own immune memory. In technical terms, it is a fail-safe mechanism that ensures the system only activates when it encounters the correct “address” in the genomic sequence.
Designing the Future: Software Tools and Algorithmic Optimization
Because the effectiveness of a gene edit depends entirely on the sequence of the sgRNA, its design has become a major focus of bioinformatics and computational biology. Designing an sgRNA is no longer a manual process; it is a task managed by sophisticated algorithms and cloud-based platforms.
Computational Challenges in sgRNA Design
The greatest technical challenge in using sgRNA is avoiding “off-target” effects. This occurs when the sgRNA binds to a DNA sequence that is similar, but not identical, to the target. In a clinical or industrial setting, an off-target cut could lead to unintended mutations or cellular dysfunction. To mitigate this, developers use computational tools like Benchling, CHOPCHOP, and CRISPR-ERA to simulate how a specific sgRNA sequence will interact with the entire genome. These tools calculate “specificity scores” based on thermodynamic stability and sequence homology.
AI and Machine Learning in Predicting On-Target Efficiency
Recent advancements in artificial intelligence have taken sgRNA design to the next level. Deep learning models are now trained on massive datasets of successful and failed CRISPR experiments to predict “on-target efficiency.” These AI tools can analyze subtle factors—such as the secondary structure of the RNA, the GC content (the ratio of guanine to cytosine), and the chromatin accessibility of the target site—that a human researcher might overlook. By applying machine learning, scientists can now design sgRNAs that are not only precise but also highly potent, ensuring a high rate of successful edits in the first attempt.
Beyond Gene Knockouts: Emerging Technical Variations of sgRNA
As the technology has matured, the role of sgRNA has expanded. It is no longer just about cutting DNA; it is about providing a programmable platform for a variety of genomic manipulations. This has led to the development of “modified” sgRNAs that serve different technical purposes.
dCas9 and CRISPR Activation/Interference (CRISPRa/i)
Engineers have developed a “dead” Cas9 (dCas9) that has lost its ability to cut DNA but retains its ability to be guided by sgRNA. In this setup, the sgRNA acts as a delivery vehicle for various “payloads,” such as transcriptional activators or repressors. This allows scientists to turn specific genes on or off without actually altering the DNA sequence—a process known as epigenetic modulation. This technical shift transforms sgRNA from a tool for “editing” into a tool for “regulation,” akin to adjusting the volume on a speaker rather than rewiring the circuit.
Multiplexing and High-Throughput Screening
One of the most powerful technical advantages of sgRNA is its scalability. Researchers can introduce a library containing thousands of different sgRNAs into a population of cells simultaneously. This is known as multiplexing. In a single experiment, a laboratory can test the function of every gene in a genome to see which ones are responsible for drug resistance or viral infection. This high-throughput approach is fundamental to modern functional genomics and is driving the discovery of new therapeutic targets at an exponential pace.
The Impact on Digital Biology and Therapeutics
The refinement of sgRNA technology is bridging the gap between digital data and physical biological systems. We are moving toward a future where “Bio-IT” is a integrated sector, and sgRNA is the fundamental interface.
Data-Driven Medicine and the Cloud
The workflow of modern gene editing often begins in the cloud. A researcher identifies a genetic mutation in a patient’s sequencing data, uses an online algorithm to design an optimized sgRNA, and orders the synthetic molecule from a manufacturer via an API. The sgRNA is then synthesized using automated chemical processes and delivered to the lab. This seamless integration of digital information and molecular manufacturing is the hallmark of Fourth Industrial Revolution technologies.
The Rise of Programmable Therapeutics
Looking forward, sgRNA represents the pinnacle of “programmable medicine.” Unlike traditional small-molecule drugs that interact with proteins in a somewhat non-specific way, sgRNA-based therapies are designed to address the root cause of genetic disease at the code level. As delivery technologies (like lipid nanoparticles) improve, the ability to deploy sgRNAs directly into human cells will enable real-time “patching” of genetic defects. This is the ultimate goal of the biotech industry: a world where disease is treated not with general chemicals, but with precise, guide-driven instructions that restore the body’s natural software to its optimal state.
In summary, sgRNA is far more than a simple biological molecule. It is the core targeting logic of the CRISPR system, a triumph of molecular engineering, and the bridge between computational design and genomic reality. As we continue to refine our ability to design and deploy these guides, the potential to reprogram biology for the betterment of human health and technology remains virtually limitless.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.