In the rapidly evolving landscape of the life sciences, the intersection of biology and technology has birthed a new era of discovery. At the heart of this convergence is a fundamental biochemical concept known as the isoelectric point, or pI. While traditionally relegated to textbooks and manual laboratory titration, pI has transitioned into a critical data point within the realm of high-tech bioinformatics, pharmaceutical software development, and artificial intelligence.
Understanding “what is pi in biochemistry” today requires looking through the lens of digital transformation. It is no longer just a measure of pH; it is a foundational variable in the algorithms that power drug discovery, protein engineering, and personalized medicine. As we move deeper into the decade of biotechnology, the technological infrastructure surrounding pI calculation and application has become a cornerstone of the modern “Bio-IT” stack.
The Digital Architecture of Protein Science
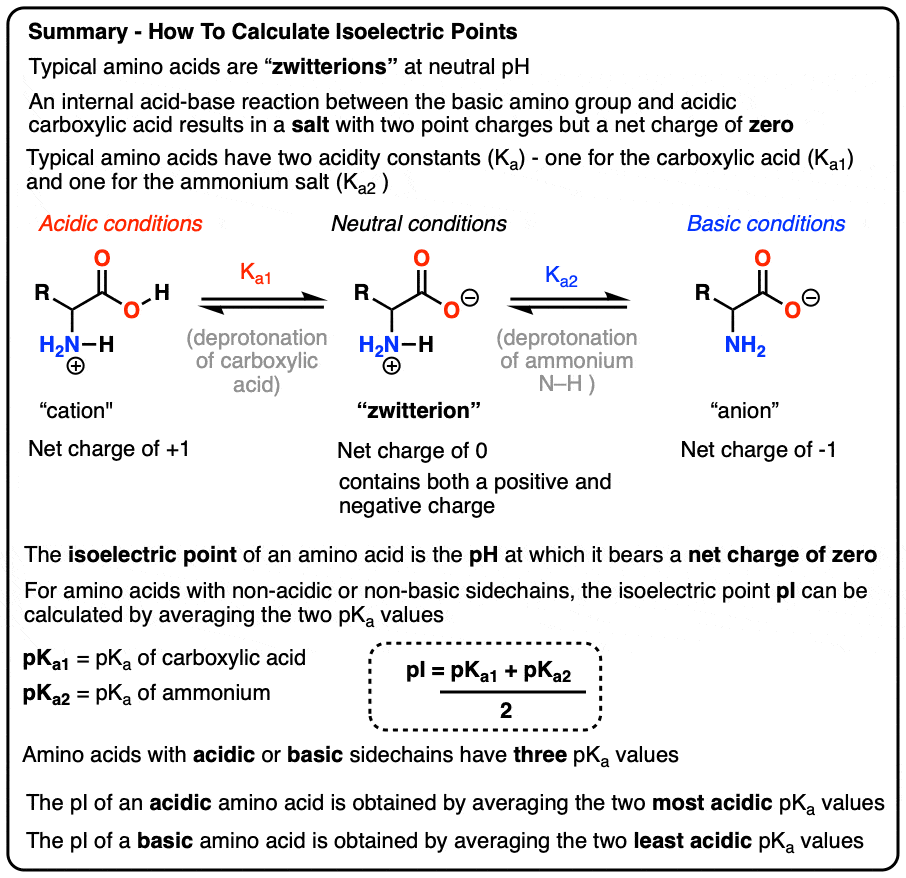
In biochemistry, the isoelectric point (pI) is the specific pH at which a molecule, typically a protein or amino acid, carries no net electrical charge. In a digital context, this value is a vital parameter used to categorize and manipulate biological data. The transition from physical observation to digital modeling has changed how scientists approach this variable.
Understanding pI in the Context of Data Modeling
To a software engineer or a bioinformatician, a protein is essentially a complex string of data—a sequence of amino acids with specific attributes. The pI is one of the most important “metadata” tags for these sequences. In computational biology, the pI dictates how a protein model will behave in a simulated environment.
When developers build databases like UniProt or Swiss-Prot, the pI is calculated using sophisticated algorithms that account for the pKa values of individual amino acids. These digital models allow researchers to predict how a protein will interact with membranes, other proteins, or synthetic drugs without ever stepping into a “wet lab.” This predictive modeling is the bedrock of the modern tech-driven approach to biochemistry.
Computational Tools for Predicting Isoelectric Points
Historically, calculating the pI was a laborious manual process. Today, a suite of software tools and web-based applications has automated this task. Tools such as ExPASy’s Compute pI/Mw tool have become industry standards. These applications use curated datasets and refined mathematical models to provide instant results.
The technology behind these tools involves parsing FASTA sequences (a text-based format for representing nucleotide or peptide sequences) and applying iterative loops to solve the Henderson-Hasselbalch equation across the entire length of a protein. This automation is not just about speed; it is about the ability to process “Big Data.” In the era of proteomics, where scientists analyze thousands of proteins simultaneously, these tech tools are indispensable.
AI and Machine Learning: Enhancing Accuracy in Biochemical Analysis
While basic algorithms can estimate pI, the biological reality is often more complex. A protein’s environment, its three-dimensional folding, and post-translational modifications can all shift its actual isoelectric point. This is where Artificial Intelligence (AI) and Machine Learning (ML) enter the frame, pushing the boundaries of what is possible in biochemical tech.
Moving Beyond Theoretical pI: The Role of Neural Networks
Standard software often calculates a “theoretical pI” based on a linear sequence of amino acids. However, in the tech world, “good enough” is being replaced by “hyper-accurate.” Advanced neural networks are now being trained on massive datasets of experimentally determined pI values. These AI models can recognize patterns that the human eye—and traditional math—might miss.
For instance, machine learning models can account for “neighboring group effects,” where the proximity of one amino acid to another changes its electrical properties. By using deep learning, tech firms are creating “Digital Twins” of proteins. These simulations provide a more nuanced understanding of how a protein will behave in the human body, significantly reducing the failure rate in digital drug trials.
Deep Learning Models for Proteome-Wide Analysis
The real power of AI in this niche lies in proteome-wide analysis. Modern tech platforms can now predict the pI of every protein expressed by a genome in minutes. This is critical for “High-Throughput Screening” (HTS), a process used in the tech-heavy pharmaceutical industry to identify potential drug candidates.
By integrating AI-driven pI predictions into their workflows, biotech companies can filter through millions of compounds to find the ones with the physical properties necessary for stability and efficacy. This synergy between biochemical theory and AI processing power is drastically shortening the timeline for bringing life-saving tech to market.
The Tech Stack of Modern Laboratories
The laboratory of 2024 is more of a data center than a traditional chemistry room. The hardware used to measure and utilize the pI of molecules is deeply integrated with sophisticated software ecosystems.
Liquid Chromatography and Mass Spectrometry Software
One of the primary ways pI is used in industry is through Isoelectric Focusing (IEF), a technique that separates proteins based on their pI. Modern IEF is often coupled with Mass Spectrometry (MS) and Liquid Chromatography (LC). The “tech stack” here includes high-end sensors and proprietary software that translates physical separation into digital peaks and graphs.
Companies like Thermo Fisher Scientific and Agilent Technologies provide integrated software suites that manage the entire data lifecycle. These platforms use the pI to orchestrate the separation process, ensuring that the hardware operates with nanometer-level precision. The integration of cloud computing allows this data to be uploaded, analyzed, and shared globally in real-time, fostering international collaboration in the tech-bio space.
Cloud-Based Platforms for Collaborative Research
The shift to the cloud has been a game-changer for biochemistry. Platforms like Benchling or AWS for Health provide a unified environment where biochemical data, including pI values, can be stored and analyzed using scalable compute power.
These platforms offer “Electronic Lab Notebooks” (ELNs) that automatically pull pI calculations from sequence data, ensuring consistency across large research teams. This digital infrastructure is essential for the “DevOps” of biology, where the goal is to iterate on protein designs with the same speed and rigor that a software company iterates on code.
Digital Security and Ethical Considerations in Bio-Tech
As biochemistry becomes increasingly digitized, the data surrounding pI and protein structures becomes highly valuable intellectual property (IP). This brings the conversation to the critical realm of digital security.
Protecting IP in Automated Proteomics
The sequences and their associated pI data are the blueprints for the next generation of biologics and vaccines. In the wrong hands, this data could be used for corporate espionage or even the creation of bio-threats. Consequently, cybersecurity is now a top priority for biotech firms.
Protecting the “pI data” involves advanced encryption, multi-factor authentication for lab equipment, and secure API integrations between different pieces of biochemical software. Tech-savvy biochemistry firms are now hiring Chief Information Security Officers (CISOs) to oversee the protection of their digital biological assets.
Data Integrity in Computational Biochemistry
Beyond security, there is the issue of data integrity. In a tech-driven environment, an error in an algorithm’s pI calculation could lead to a cascade of failures in drug development. Ensuring the “provenance” of data—knowing exactly where a pI value came from and how it was calculated—is essential.
Blockchain technology is beginning to see exploration in this area as a way to create an immutable audit trail for biochemical research. By logging every calculation and modification on a decentralized ledger, the tech community can ensure that the foundational data used in drug discovery remains untampered and verifiable.
The Future: Quantum Computing and the Next Frontier of pI
As we look toward the future, the limits of classical computing are becoming apparent in the face of biological complexity. The next major leap in biochemistry tech will likely come from Quantum Computing.
Quantum computers excel at simulating molecular interactions at an atomic level—something that is incredibly difficult for binary systems. In the context of the isoelectric point, a quantum computer could theoretically calculate the “absolute pI” of a protein in a dynamic, changing environment with 100% accuracy.
This would represent the ultimate convergence of tech and biochemistry. We would move from “predicting” protein behavior to “knowing” it. As tech giants like Google and IBM continue to advance quantum hardware, the biochemical world waits on the cusp of a revolution where the pI is no longer a static number, but a dynamic, digitally mastered variable in the quest to solve the mysteries of life.
Conclusion: The Synthesis of Code and Carbon
The question of “what is pi in biochemistry” is no longer a simple one. In the modern world, it is a bridge between the biological world of carbon and the technological world of silicon. From the basic algorithms used in bioinformatics to the cutting-edge neural networks of AI, the isoelectric point remains a vital metric.
As technology continues to advance, our ability to calculate, manipulate, and secure this biochemical data will define the future of medicine, industry, and scientific understanding. In this high-tech era, the biochemist and the software engineer are working on two sides of the same coin, using the pI as a common language to build a healthier, more advanced future.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.