In the rapidly evolving world of technology, we often look to silicon and software for the next great breakthrough. However, the most sophisticated operating system ever designed isn’t running on a server in Silicon Valley; it is running within the nucleus of every living cell. As we move deeper into the era of biotechnology and high-performance computing, the lines between computer science and genetics have blurred. To understand the future of health tech, personalized medicine, and synthetic biology, one must understand the fundamental architecture of our genetic “source code.” At the heart of this architecture are two critical components: exons and introns.
While these terms originate from molecular biology, they represent a complex system of data management, modular programming, and signal processing that mirrors the most advanced software engineering principles. This article explores the technical nuances of exons and introns and examines how their understanding is driving the next wave of technological innovation in bioinformatics and artificial intelligence.
Decoding the Software of Life: What Are Exons and Introns?
To understand exons and introns, we must first view deoxyribonucleic acid (DNA) as a massive, encrypted database. This database contains the instructions for building and operating an organism. However, unlike a simple text file, DNA is structured with a sophisticated “cut-and-paste” logic.
The Anatomy of a Gene
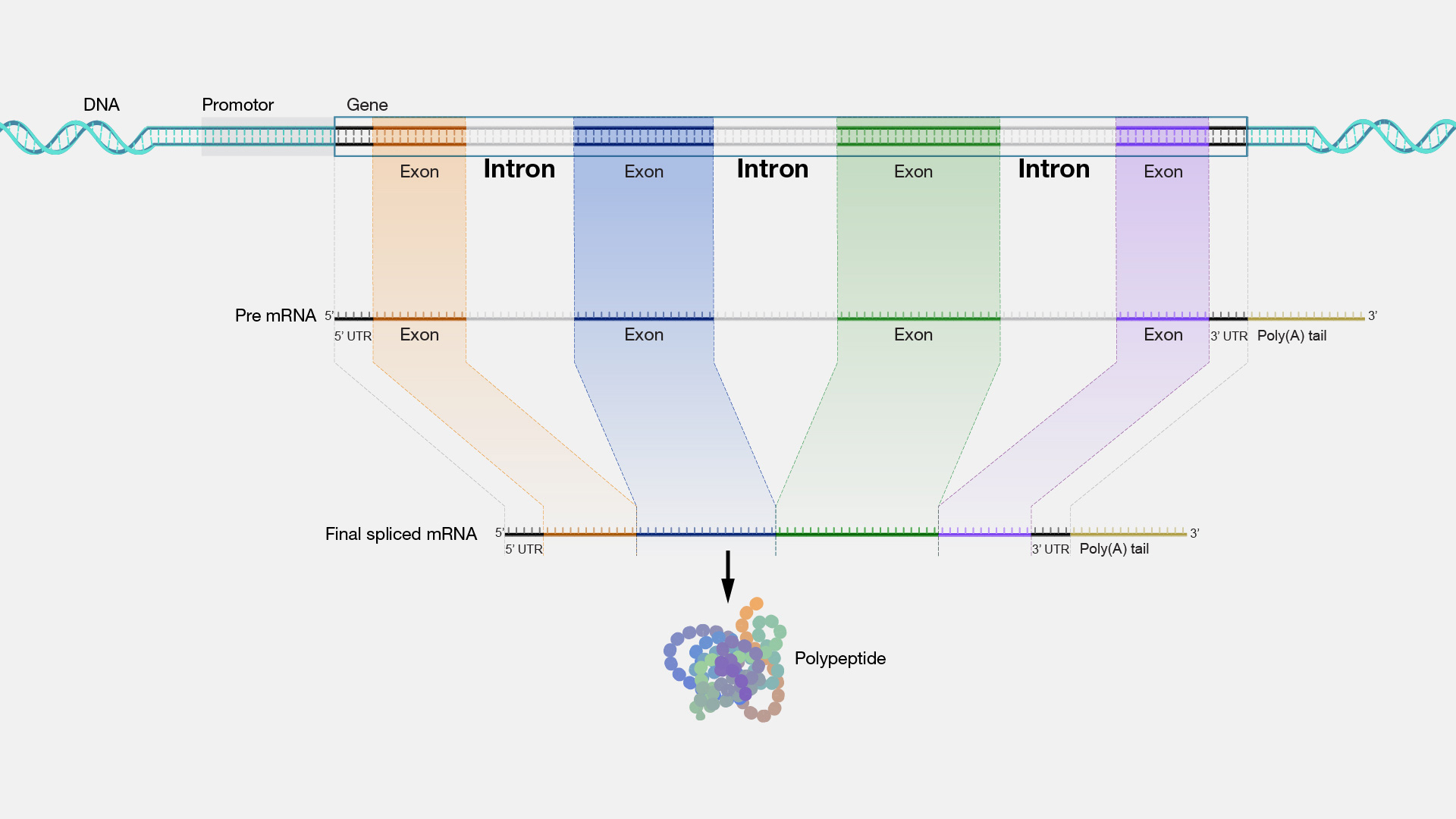
In the context of genomic data, a gene is a functional unit of inheritance. When a cell needs to produce a protein—the physical machinery of life—it doesn’t just read the DNA linearly and produce a finished product. Instead, the gene undergoes a process called transcription. The resulting molecule, pre-messenger RNA (pre-mRNA), contains both coding and non-coding sequences.
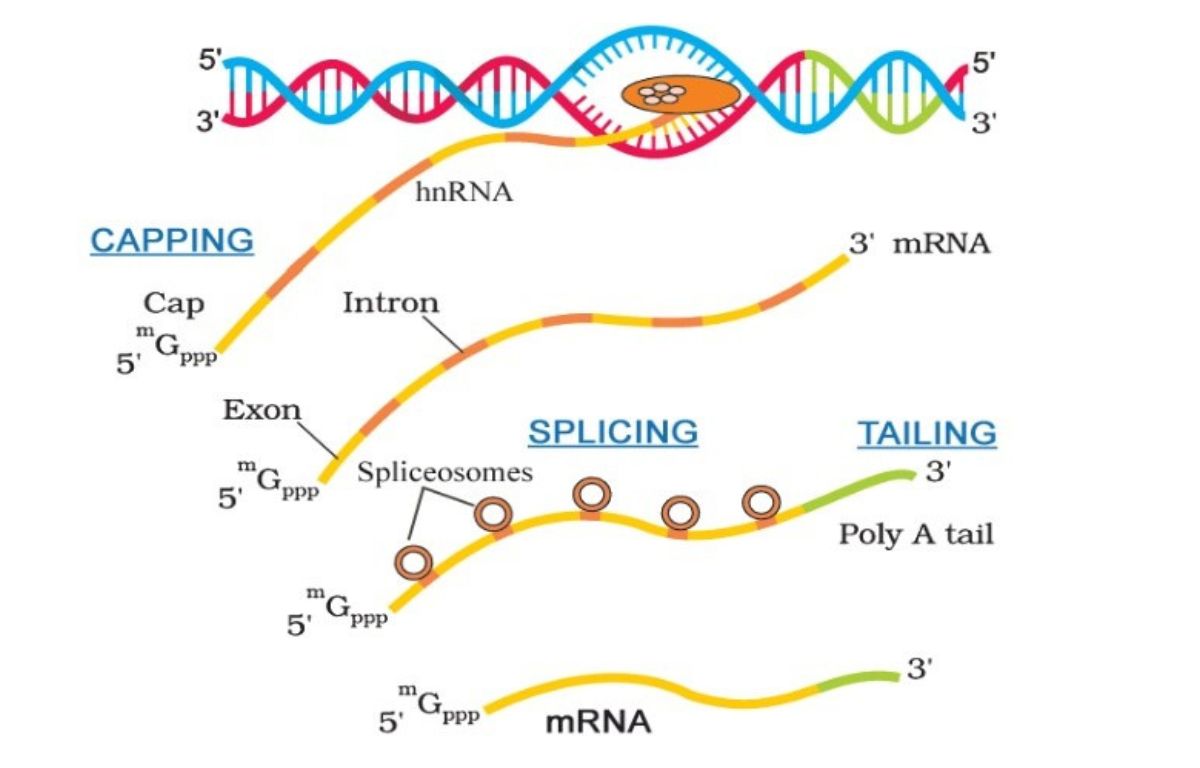
Exons (short for “expressed regions”) are the sequences of DNA that provide the actual code for proteins. They are the functional segments that carry the “instructions” for the hardware. Introns (short for “intervening regions”) are the segments located between exons. For a long time, introns were dismissed by early researchers as “junk DNA,” but we now understand them as vital regulatory elements that manage how and when the exons are utilized.
Information vs. Interruption: The Splicing Mechanism
The transition from pre-mRNA to mature mRNA involves a highly technical process known as “splicing.” Think of this as a compiler in a programming environment. The spliceosome—a complex molecular machine—identifies the introns, removes them, and stitches the exons together into a continuous, readable string of data.
This process is not merely a biological necessity; it is a masterpiece of data efficiency. Through a phenomenon called “alternative splicing,” a single gene can produce multiple different protein versions by choosing which exons to include or exclude. In tech terms, this is equivalent to a single source code repository generating multiple different software builds depending on the configuration settings.
The Technological Parallel: Why Software Engineers Should Care About Splicing
The structure of exons and introns provides a fascinating parallel to modern software architecture, particularly in the realms of modularity and data compression. As we build more complex AI models, the “exon-intron” model offers a blueprint for efficient information storage.
Data Compression and Efficiency
One of the greatest challenges in Big Data is the “signal-to-noise” ratio. In a human genome of 3 billion base pairs, only about 1.5% consists of exons (the protein-coding sequences). The rest is non-coding. From a data science perspective, this looks like a massive waste of storage. However, much like metadata in a digital file, the “non-coding” introns provide the instructions for how the coding data should be processed.
In software engineering, we often use abstraction layers. Introns act as a biological abstraction layer, allowing the “exon” data to be reused in different contexts. This modularity allows for high levels of biological complexity without needing an exponentially larger genome. Technologists are now looking at these biological “compression algorithms” to develop new ways of storing data, including DNA-based data storage, where digital files are encoded into synthetic DNA strands.
The Role of AI in Gene Sequencing
Identifying where an exon ends and an intron begins (splice site prediction) is a massive computational challenge. Traditional algorithms often struggle with the sheer variability of genetic sequences. This is where machine learning and Deep Learning have stepped in.
AI tools, such as deep neural networks, are now being trained on massive genomic datasets to predict splicing patterns with incredible accuracy. Tools like SpliceAI use residual convolutional neural networks to identify the “logic gates” of our DNA. For the tech industry, this represents a new frontier: using AI to debug the human code. By identifying mutations in introns that lead to “splicing errors,” software-driven diagnostics can catch diseases like spinal muscular atrophy or certain cancers long before symptoms appear.
Breakthroughs in Biotech: Leveraging Genetic Logic for Future Tech
The mastery of exons and introns isn’t just an academic exercise; it is the foundation of the burgeoning “Bio-IT” sector. We are moving from a phase of simply “reading” the code to actively “editing” and “writing” it.
CRISPR and the Editing Revolution
CRISPR-Cas9 is often described as a “word processor” for DNA. However, the tech is far more nuanced. To effectively edit a gene, researchers must understand the exon-intron boundaries to ensure they don’t disrupt the regulatory framework of the cell. If you delete an exon, you lose the protein; if you delete an intron, you might lose the “timer” or “switch” that tells the protein when to turn on.
Current gene therapies are being designed to target introns specifically. By using “antisense oligonucleotides” (ASOs)—essentially small snippets of code—technologists can force a cell to skip a mutated exon or include a missing one. This is the ultimate “hotfix” for a biological system, and it is entirely dependent on our digital mapping of exon-intron architecture.
Synthetic Biology and Programming Organisms
Synthetic biology is the practice of designing and constructing new biological parts, devices, and systems. In this field, exons are treated as “standardized parts” or “bio-bricks.” Engineers are building “gene circuits” where exons are swapped in and out to create organisms that can perform specific tasks, such as secreting insulin or breaking down plastic in the ocean.
This modular approach is identical to object-oriented programming (OOP). By treating exons as objects and introns as the logic that dictates their interaction, synthetic biologists are creating a new form of “living software.” The potential for this tech is limitless, ranging from “living sensors” that detect toxins to “bio-computers” that can store data and perform logic operations within a cell.
Data Security and Bio-Cryptography: Protecting the “Exon” Assets
As our genetic information becomes increasingly digitized, the security of this data becomes a paramount tech concern. Your “exon sequence” is the ultimate biometric—it is the source code of you.
The Vulnerability of Genetic Data
With the rise of consumer genomics and digital health platforms, millions of people’s exon data are stored on cloud servers. This has created a new category of cybersecurity: bio-security. Unlike a password or a credit card number, you cannot change your DNA if it is leaked. The technical challenge lies in “de-identifying” genomic data while still making it useful for researchers.
Introns, which vary significantly between individuals, are often used as “genetic fingerprints.” Tech firms are now developing encrypted computational environments where researchers can analyze the “exons” (the functional data) without ever having access to the “introns” (the personal identifiers), ensuring privacy through mathematical certainty.
Blockchain in Genomic Security
To combat the risk of data breaches, some tech startups are looking to blockchain technology. By storing genomic data on a decentralized ledger, individuals can maintain ownership of their “exon-intron” sequences. Using smart contracts, a user can grant a pharmaceutical company temporary access to their data for a specific study, receiving payment in return, all while the data remains encrypted and sovereign. This represents a radical shift in how we value biological information as a digital asset.
The Future of Digital Health: Predictive Algorithms and Genetic Splicing
The final frontier of this technological journey is the move toward predictive, personalized medicine. We are entering an era where your “digital twin”—a computer model of your biology—can simulate how your specific exon-intron structure will react to a new drug.
Advanced software platforms are now capable of “in silico” testing. Instead of a patient taking a drug to see if it works, a cloud-based algorithm simulates the drug’s effect on the patient’s specific splice variants. This reduces the risk of adverse reactions and increases the efficacy of treatments.
As we look forward, the integration of AI, cloud computing, and molecular biology will continue to accelerate. The study of exons and introns has moved out of the laboratory and into the data center. By understanding the “logic of life,” tech leaders are not just building better apps or faster processors; they are unlocking the secrets of longevity, health, and the very essence of what it means to be human. In the binary of the future, the most important digits won’t just be 0 and 1, but A, C, T, and G—and the complex ways we splice them together.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.