In the rapidly evolving landscape of biotechnology, the ability to read, interpret, and manipulate the human genome has transitioned from science fiction to a standard of modern medicine. At the heart of this revolution lies the study of structural variations within our DNA. One of the most complex phenomena in this field is the “balanced translocation.” While traditionally viewed through a purely biological lens, understanding balanced translocation in the modern era requires a deep dive into high-throughput sequencing, bioinformatics, and the sophisticated hardware that allows us to map the invisible architecture of our cells.

The Bio-Technical Architecture of Balanced Translocation
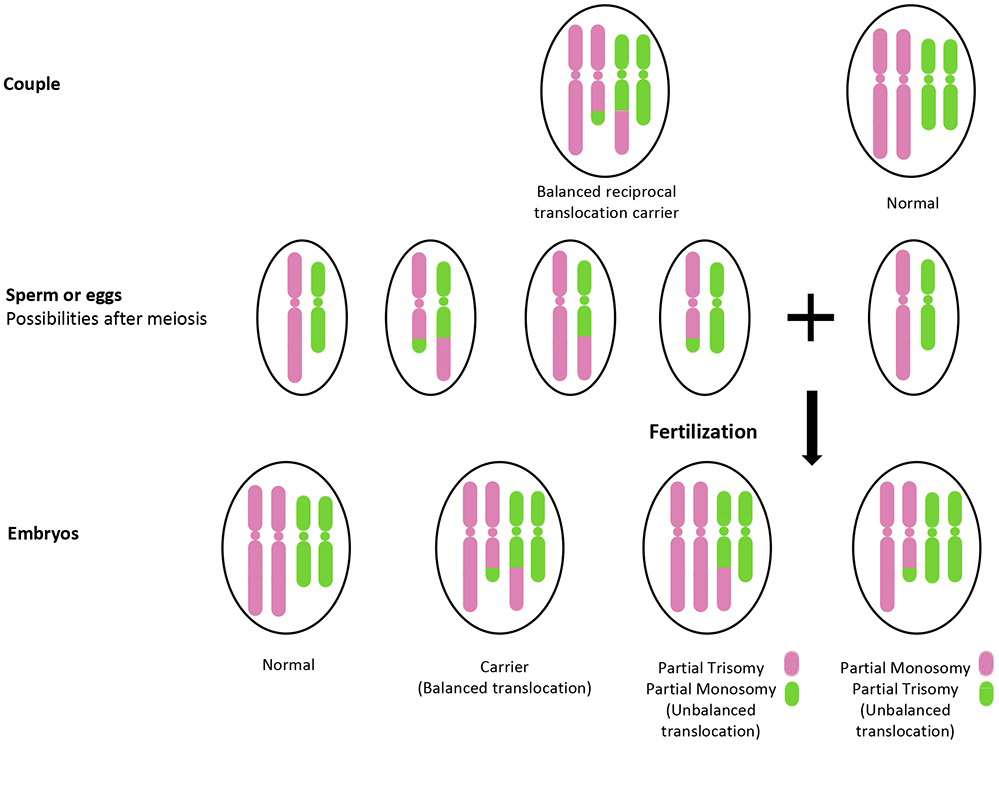
To understand balanced translocation from a technical perspective, one must view the human genome as a massive database of instructions organized into 23 pairs of chromosomal “folders.” A balanced translocation occurs when segments of two chromosomes break off and switch places. The term “balanced” is critical: it implies that no genetic material has been lost or gained during this swap. The individual remains functionally healthy because the complete “software code” of their body is present, even if it is stored in the wrong location.
Understanding the “Switch”: How Chromosomal Rearrangements Occur
From a systems engineering standpoint, a balanced translocation is a structural bug that doesn’t necessarily crash the program. During the process of meiosis or early development, physical stress or chemical triggers can cause a double-strand break in the DNA helix. When the cellular repair mechanisms—essentially the body’s automated error-correction protocols—attempt to fix these breaks, they occasionally misroute the segments. If Chromosome 4 and Chromosome 20 swap pieces perfectly, the “system” continues to run smoothly. However, the technical challenge arises during reproduction, where these “misplaced folders” can lead to unbalanced offspring, necessitating high-tech intervention.
The Digital Mapping of Genetic Material
The identification of these translocations relies on our ability to visualize the physical structure of chromosomes. In the early days of tech, this was done via basic microscopy (karyotyping). Today, we use digital imaging and automated pattern recognition software to create a high-resolution “map” of the genome. By digitizing the chromosomal landscape, technicians can identify exactly where a break has occurred, down to the specific base pair. This digital mapping is the foundation upon which all subsequent genetic counseling and intervention are built.
Advanced Diagnostic Tools: From Microarrays to Next-Generation Sequencing (NGS)
Identifying a balanced translocation is notoriously difficult because standard diagnostic tests often look for “missing” or “extra” data. Since a balanced translocation involves no change in the quantity of DNA, older software algorithms frequently overlook it. The tech industry has responded by developing high-sensitivity diagnostic suites that prioritize structural orientation over simple volume.
Fluorescence In Situ Hybridization (FISH) in the Digital Age
FISH is a powerful imaging technology that uses fluorescent probes to “tag” specific sequences of DNA. When viewed under high-powered digital microscopes equipped with specialized sensor arrays, these tags glow in different colors. If a green tag (from Chromosome A) appears on a chromosome that should be entirely red (Chromosome B), the software flags a translocation. Modern FISH tech has been enhanced by automated scanning platforms that can process hundreds of samples simultaneously, reducing human error and increasing the throughput of clinical labs.
The Role of Comparative Genomic Hybridization (aCGH)
Array Comparative Genomic Hybridization (aCGH) represents a leap forward in the digitalization of genetics. This technology uses “chips” populated with thousands of defined DNA probes. While aCGH is excellent at detecting “unbalanced” variations (where data is missing), it has historically struggled with balanced versions. However, the newest generation of “SNP-microarrays” uses sophisticated signal-processing algorithms to detect “Loss of Heterozygosity” and other subtle markers that hint at a balanced rearrangement, providing a more comprehensive view of the patient’s genomic health.
AI and Machine Learning in Genetic Pattern Recognition
The sheer volume of data produced by a single human genome—approximately 3 billion base pairs—is far too vast for manual review. This is where Artificial Intelligence (AI) and Machine Learning (ML) have become the most critical tools in the geneticist’s arsenal.
Algorithmic Detection of Structural Variants
AI models, specifically Deep Learning architectures like Convolutional Neural Networks (CNNs), are now trained to recognize the visual signatures of balanced translocations in karyotype images. These algorithms are trained on millions of labeled genomic images, allowing them to spot deviations that the human eye might miss. By automating the initial screening process, these AI tools allow laboratories to scale their operations and provide faster results to patients.
Predictive Modeling for Reproductive Success
For individuals with a balanced translocation, the primary concern is often the risk of passing on an “unbalanced” version to their children. Tech startups are now developing predictive modeling software that simulates millions of possible meiotic outcomes. By analyzing the specific breakpoints in a parent’s translocation, these programs can calculate the exact percentage of viable vs. non-viable embryos. This data-driven approach allows for more informed decision-making in fertility treatments, such as In Vitro Fertilization (IVF).
The Integration of Software Solutions in Clinical Labs
The transition from “wet lab” biology to “dry lab” data analysis has necessitated a robust software infrastructure. Managing the data of patients with balanced translocations requires highly secure, scalable, and interoperable software systems.
Laboratory Information Management Systems (LIMS)
Modern genetics labs run on LIMS—complex software ecosystems that track a sample from the moment a swab is taken to the final report generation. For balanced translocation cases, LIMS must integrate data from various sources: imaging from FISH, raw data from NGS, and clinical notes from genetic counselors. The goal is to create a “Single Source of Truth” for the patient’s genomic data, ensuring that no technical nuance is lost in translation between different medical departments.
Data Privacy and the Security of Genomic Clouds
Genomic data is the most sensitive information a person possesses. As labs move their processing to the cloud to take advantage of high-performance computing (HPC) for sequencing, cybersecurity becomes paramount. Encryption protocols (such as AES-256) and decentralized data storage are being implemented to ensure that a person’s structural genetic “map” is never leaked or misused. In the era of personalized medicine, protecting the digital integrity of the genome is as important as the medical treatment itself.
The Road Ahead: CRISPR and the Potential for Technical Correction
If we view a balanced translocation as a “misplaced line of code,” the ultimate technological goal is to move that code back to its original location. This brings us to the cutting edge of tech: CRISPR-Cas9 and other gene-editing platforms.
Gene Editing as the Ultimate Software Patch
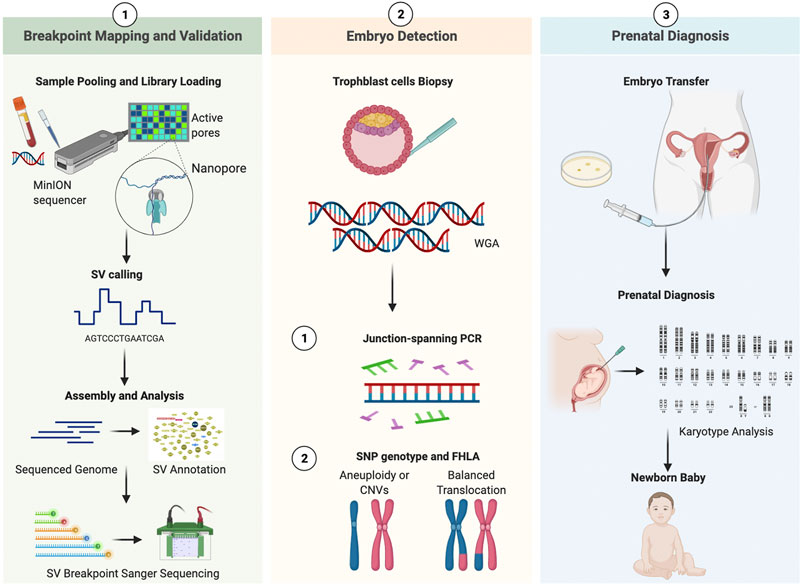
CRISPR acts as a biological “search and replace” tool. While we are currently in the early stages of using this tech for structural rearrangements, the potential is staggering. Researchers are working on “Prime Editing” and “Base Editing” technologies that could, in theory, target the breakpoints of a balanced translocation and swap the segments back to their native chromosomes. This would effectively “patch” the genome, eliminating the risk of unbalanced offspring.
The Ethics of Technical Intervention
As with any disruptive technology, the ability to “re-map” the human genome brings significant ethical considerations. The tech community is currently debating the governance of these tools. Should we only correct translocations that cause disease, or do we move toward a future of “genomic optimization”? As our hardware becomes more precise and our AI models more predictive, the line between medical necessity and technical enhancement will continue to blur.
In conclusion, a balanced translocation is more than just a biological curiosity; it is a complex structural challenge that is being solved through the marriage of biology and technology. From the high-resolution sensors of modern microscopes to the deep learning algorithms that process genomic data, technology is providing the clarity needed to navigate our own DNA. As we continue to refine these tools, the dream of perfectly understanding—and perhaps one day correcting—the architectural quirks of the human genome moves closer to reality.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.