Chromatin, the intricate complex of DNA and proteins that forms chromosomes within the nucleus of eukaryotic cells, is far more than just a passive packaging mechanism. It’s a dynamic regulatory hub, a sophisticated scaffold that dictates gene expression, DNA replication, and repair. For decades, understanding this fundamental biological structure was largely confined to biochemical assays and microscopy. However, in recent years, a revolution in technology has transformed chromatin from a subject of inference to one of direct, high-resolution interrogation. This technological leap has opened unprecedented avenues for dissecting the complexities of our genome, with profound implications for drug discovery, diagnostics, and fundamental biological understanding.
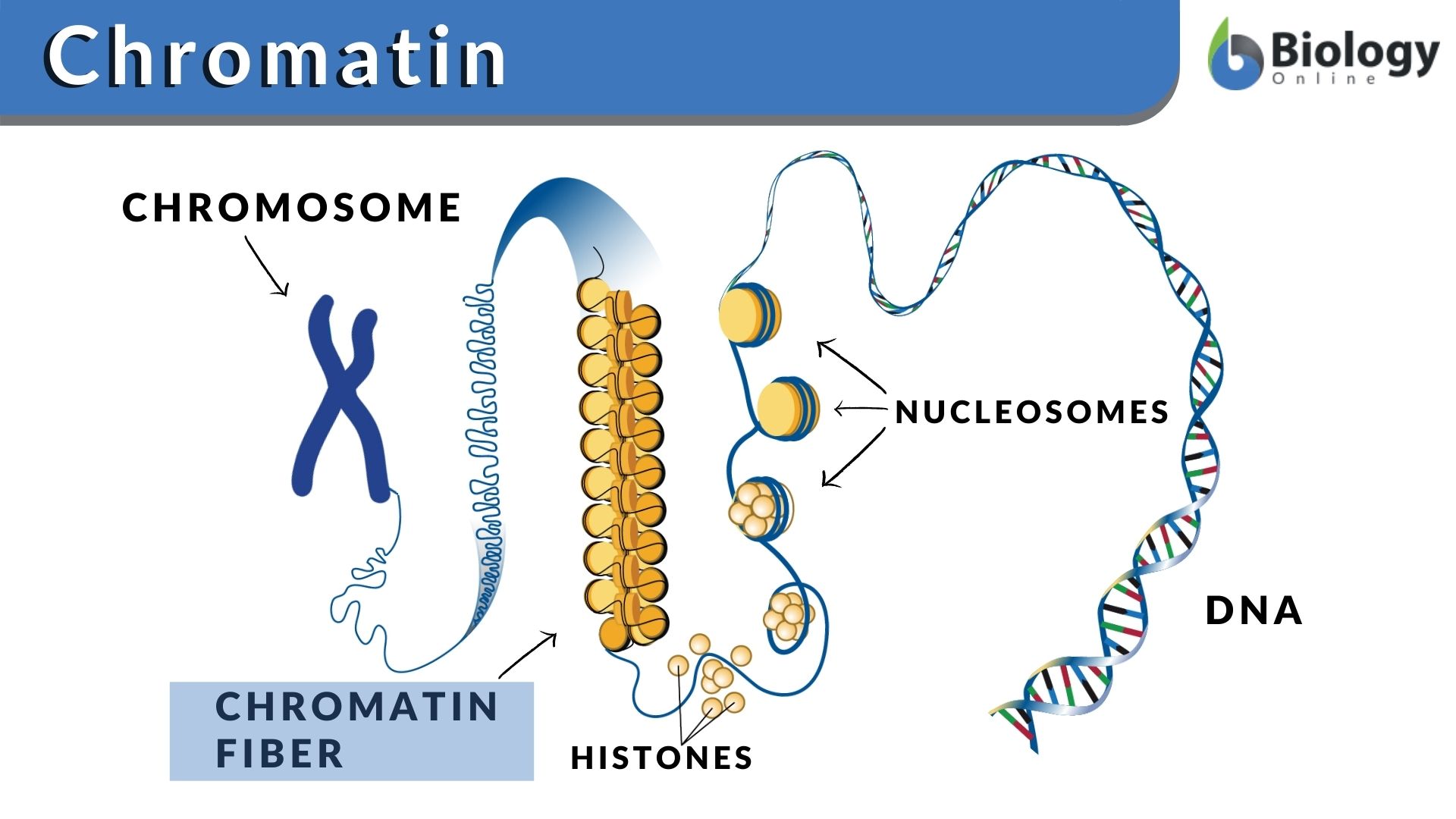
The “what” of chromatin in a technological context refers to the suite of advanced tools and methodologies that allow us to visualize, map, manipulate, and analyze its structure and function at an ever-increasing scale and resolution. It’s about the innovations that enable us to move beyond static snapshots and witness the dynamic dance of DNA, histones, and regulatory proteins. This article delves into the technological landscape that has brought chromatin research to the forefront of biological discovery, exploring the key innovations, their applications, and the future directions driven by relentless technological advancement.
H2: Illuminating Chromatin: The Rise of High-Throughput Sequencing and Epigenomic Mapping
The advent of Next-Generation Sequencing (NGS) technologies has been a cornerstone in unraveling the organizational principles of chromatin. Before NGS, studying epigenetic modifications and chromatin accessibility was a laborious, low-throughput process. Now, we can map these features across entire genomes with remarkable speed and cost-effectiveness. These technologies have empowered researchers to ask broader and more complex questions about how chromatin structure influences cellular identity and function.
H3: Chromatin Immunoprecipitation Sequencing (ChIP-seq): Mapping Protein Interactions
ChIP-seq is arguably one of the most powerful technologies for understanding chromatin. It allows researchers to identify the genomic regions where specific proteins, such as transcription factors or histone modifications, bind. The process involves crosslinking proteins to DNA in living cells, fragmenting the chromatin, and then using an antibody specific to the protein of interest to immunoprecipitate the DNA-protein complexes. The precipitated DNA is then sequenced using NGS. The resulting sequence reads, when mapped back to the genome, reveal the precise locations of the target protein’s binding sites.
- Technological Advancements: Early ChIP methods were plagued by low sensitivity and high background noise. However, improvements in antibody quality, chromatin fragmentation techniques (e.g., sonication and enzymatic digestion), and library preparation for sequencing have dramatically increased the resolution and reliability of ChIP-seq. The development of specialized protocols for low-input samples has also been crucial, allowing ChIP-seq to be performed on rare cell populations or clinical samples where material is limited.
- Applications in Tech: In a tech context, ChIP-seq data is crucial for building comprehensive regulatory network maps. This information feeds into AI algorithms designed to predict gene expression patterns, identify potential drug targets for diseases like cancer, and understand the mechanisms of drug resistance. The sheer volume of data generated by ChIP-seq necessitates advanced bioinformatic pipelines and cloud computing infrastructure for storage, processing, and analysis.
H3: ATAC-seq and DNase-seq: Probing Chromatin Accessibility
Beyond identifying specific protein binding sites, understanding which regions of chromatin are open and accessible to the transcriptional machinery is paramount. Technologies like Assay for Transposase-Accessible Chromatin using sequencing (ATAC-seq) and DNase I hypersensitive sites sequencing (DNase-seq) serve this purpose. These methods exploit the principle that open chromatin regions are more susceptible to enzymatic cleavage.
- ATAC-seq: This method uses a hyperactive Tn5 transposase that preferentially inserts sequencing adapters into open chromatin regions. This elegantly simplifies the workflow, requiring minimal starting material and fewer steps compared to older methods. Its efficiency has made it a go-to technique for rapid profiling of chromatin accessibility.
- DNase-seq: This technique uses the enzyme DNase I to digest DNA. Regions of open chromatin are protected from degradation, while tightly packed heterochromatin is more susceptible. The resulting DNA fragments are then sequenced.
- Technological Synergy: Both ATAC-seq and DNase-seq provide complementary insights into the dynamic nature of the genome. When combined with other epigenetic data, such as ChIP-seq for histone modifications, they create a multi-layered view of gene regulation. The development of advanced peak-calling algorithms and visualization tools is essential for extracting meaningful biological insights from the vast datasets generated by these assays.
H2: Visualizing the Unseen: Advanced Imaging and Microscopy Techniques
While sequencing technologies provide high-throughput genomic information, microscopy techniques offer invaluable spatial and temporal insights into chromatin organization. The ability to visualize chromatin within living cells or in intricate detail in fixed samples has been revolutionized by innovations in microscopy, allowing us to see the physical architecture that underlies gene regulation.
H3: Super-Resolution Microscopy: Breaking the Diffraction Limit
Traditional light microscopy is limited by the diffraction of light, preventing the visualization of structures smaller than approximately 200 nanometers. Super-resolution microscopy techniques, such as STORM (Stochastic Optical Reconstruction Microscopy) and PALM (Photoactivated Localization Microscopy), have overcome this limitation. These methods utilize fluorescent molecules that can be precisely activated and localized, allowing for the reconstruction of images with resolutions down to tens of nanometers.
- Technological Underpinnings: The development of novel fluorescent probes, advanced imaging algorithms, and highly sensitive cameras are key to the success of super-resolution microscopy. These techniques allow researchers to visualize individual nucleosomes, protein complexes within chromatin fibers, and the dynamic movement of chromatin domains in real-time.
- Impact on Chromatin Research: Super-resolution microscopy provides direct visual evidence of how chromatin is folded, how nuclear organization changes during cellular processes like mitosis, and how specific proteins interact within the crowded nuclear environment. This has been instrumental in validating models derived from sequencing data and generating new hypotheses about chromatin dynamics.
H3: Cryo-Electron Microscopy (Cryo-EM): Unveiling Molecular Architecture
For visualizing the detailed 3D structures of chromatin components and their assemblies, Cryo-EM has emerged as a transformative technology. This technique involves flash-freezing a sample in a thin layer of vitreous ice, preserving its native structure, and then imaging it with an electron microscope. By collecting images of many individual molecules from different angles and computationally reconstructing them, researchers can achieve near-atomic resolution.
- Advancements in Data Processing: While electron microscopy itself is a mature technology, significant advancements in detector technology, robotic sample handling, and sophisticated image processing software have made Cryo-EM accessible and powerful for biological applications. The ability to analyze large datasets and reconstruct complex molecular machines is a testament to the computational power driving this field.
- Decoding Chromatin Complexes: Cryo-EM has provided unprecedented insights into the structure of the nucleosome, the fundamental repeating unit of chromatin, as well as the architecture of larger chromatin-modifying complexes. This molecular-level understanding is crucial for designing targeted drugs that interact with specific components of the chromatin machinery.
H2: Engineering Chromatin: Tools for Manipulation and Perturbation
Beyond observation, technological advancements now allow us to actively manipulate chromatin to study its function. Gene editing technologies, particularly CRISPR-based systems, have revolutionized our ability to alter chromatin structure and assess the consequences on gene expression and cellular phenotypes.
H3: CRISPR-based Technologies for Chromatin Engineering
The CRISPR-Cas9 system, originally discovered as a bacterial immune mechanism, has been ingeniously adapted for genome editing and, more recently, for precise manipulation of chromatin. By fusing catalytically inactive Cas9 (dCas9) to effector domains (e.g., transcriptional activators, repressors, or epigenetic modifiers), researchers can target specific genomic loci and alter their chromatin state.
- CRISPRi and CRISPRa: CRISPR interference (CRISPRi) uses a dCas9 fused to a repressor domain to block transcription by sterically hindering RNA polymerase access or by recruiting repressive chromatin modifiers. Conversely, CRISPR activation (CRISPRa) uses dCas9 fused to transcriptional activators to recruit co-activators and alter chromatin to promote gene expression.
- Epigenome Editing: Further innovation has led to the development of “epigenome editors,” which combine dCas9 with enzymes that can directly modify epigenetic marks, such as DNA methylation or histone acetylation, without altering the underlying DNA sequence. This allows for precise studies of the functional roles of specific epigenetic modifications in controlling gene expression and chromatin structure.
- Technological Scalability: The ability to design guide RNAs for multiple targets simultaneously, coupled with high-throughput screening platforms, allows for large-scale experiments to map the functional landscape of the genome. This is crucial for understanding how combinations of epigenetic marks contribute to cellular states and disease pathogenesis.
H3: Chemical Genomics and Probe Development
The development of small molecules and chemical probes that specifically inhibit or activate proteins involved in chromatin regulation is another critical technological avenue. These tools allow researchers to perturb chromatin dynamics pharmacologically and study the downstream effects.
- Targeting Epigenetic Enzymes: Inhibitors of histone deacetylases (HDACs), histone methyltransferases (HMTs), and DNA methyltransferases (DNMTs) are prime examples. These drugs have already found applications in cancer therapy, highlighting the direct translational potential of chromatin-targeted technologies.
- High-Throughput Screening: Advancements in automated screening platforms and the development of innovative assay designs allow for the rapid identification and optimization of novel chromatin-modulating compounds. This accelerates the drug discovery pipeline and provides new tools for dissecting epigenetic pathways.
H2: The Future of Chromatin Research: AI, Big Data, and Personalized Medicine
The technological evolution in chromatin research is not slowing down. The increasing volume and complexity of data generated by sequencing, imaging, and manipulation technologies necessitate the integration of artificial intelligence and advanced computational approaches.
H3: AI and Machine Learning in Chromatin Analysis
AI and machine learning algorithms are becoming indispensable for making sense of the massive datasets produced in chromatin research. These tools can identify complex patterns, predict gene regulatory networks, and even discover novel epigenetic mechanisms that would be missed by traditional analysis methods.
- Predictive Modeling: AI can be used to predict the functional impact of specific chromatin states on gene expression, identify potential disease-associated epigenetic variations, and forecast cellular responses to therapeutic interventions.
- Image Analysis: Deep learning algorithms are revolutionizing the analysis of microscopy images, enabling automated cell segmentation, feature extraction, and quantification of chromatin dynamics in complex biological samples.
- Data Integration: The ability of AI to integrate diverse datasets – including genomics, epigenomics, transcriptomics, and proteomics – offers a holistic view of cellular function and allows for the discovery of emergent properties of chromatin regulation.
H3: Toward Personalized Epigenomic Medicine
The ultimate technological frontier for chromatin research lies in its application to personalized medicine. Understanding an individual’s unique epigenomic landscape – how their chromatin is organized and regulated – holds the key to tailoring treatments for diseases.
- Diagnostic Biomarkers: Epigenetic modifications can serve as sensitive biomarkers for early disease detection and prognosis. Technologies that allow for rapid and cost-effective epigenomic profiling of patient samples are crucial for this endeavor.
- Therapeutic Target Identification: By analyzing the epigenomic differences between healthy and diseased cells, researchers can identify specific chromatin-related targets for drug development, leading to more effective and less toxic therapies.
- Monitoring Treatment Response: Tracking changes in an individual’s epigenome over time can help monitor treatment response and guide therapeutic adjustments, moving towards a truly personalized and adaptive approach to healthcare.
In conclusion, the question “what does chromatin” has evolved from a purely biological inquiry to a profound technological challenge and opportunity. The sophisticated tools and methodologies discussed here have transformed our ability to investigate this fundamental aspect of cellular biology. From high-throughput sequencing and advanced microscopy to gene editing and AI-driven analysis, technology is continuously pushing the boundaries of our understanding, paving the way for groundbreaking discoveries and ultimately, for innovations that could revolutionize medicine and our fundamental comprehension of life itself.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.