In the rapidly evolving landscape of data science, bioinformatics, and statistical computing, the R programming language remains a cornerstone for researchers and developers alike. While “Base R” provides a robust foundation for mathematical operations and basic data handling, the true power of the language lies in its modularity. This modularity is facilitated through packages—collections of functions, data, and compiled code that expand R’s capabilities. Whether you are building complex neural networks, creating stunning visualizations with ggplot2, or cleaning messy datasets with tidyverse, knowing how to efficiently manage these extensions is a fundamental technical skill.
This guide provides an in-depth exploration of how to install packages in R, covering everything from the standard Comprehensive R Archive Network (CRAN) to advanced installation techniques from GitHub and source files.
The Foundation of R Packages: Understanding the Ecosystem
Before diving into the “how,” it is essential for a technology professional to understand the “where.” The R ecosystem is decentralized yet highly organized, relying on several key repositories that host thousands of open-source contributions.
What is CRAN?
The Comprehensive R Archive Network (CRAN) is the primary repository for R packages. It is a network of ftp and web servers around the world that store identical, up-to-date versions of code and documentation for R. When you install a package from CRAN, you are benefiting from a rigorous vetting process. CRAN maintainers ensure that the package passes a battery of tests across different operating systems (Windows, macOS, and Linux) to ensure stability and compatibility.
The Role of Bioconductor and GitHub
While CRAN is the general-purpose hub, other repositories serve specific niches. Bioconductor is a specialized repository for biological data analysis, providing tools for high-throughput genomic data. Because Bioconductor has its own release cycle and dependency logic, it requires a different installation approach than CRAN.
On the other end of the spectrum is GitHub. Many developers host the “development versions” of their packages on GitHub. These versions often contain the latest features and bug fixes that have not yet been pushed to CRAN. For tech-savvy users, GitHub is the gateway to the bleeding edge of R software development.
Core Methods for Installing Packages
For most users, installation happens directly within the R console or through an Integrated Development Environment (IDE) like RStudio. There are three primary ways to handle standard installations.
Using the install.packages() Function
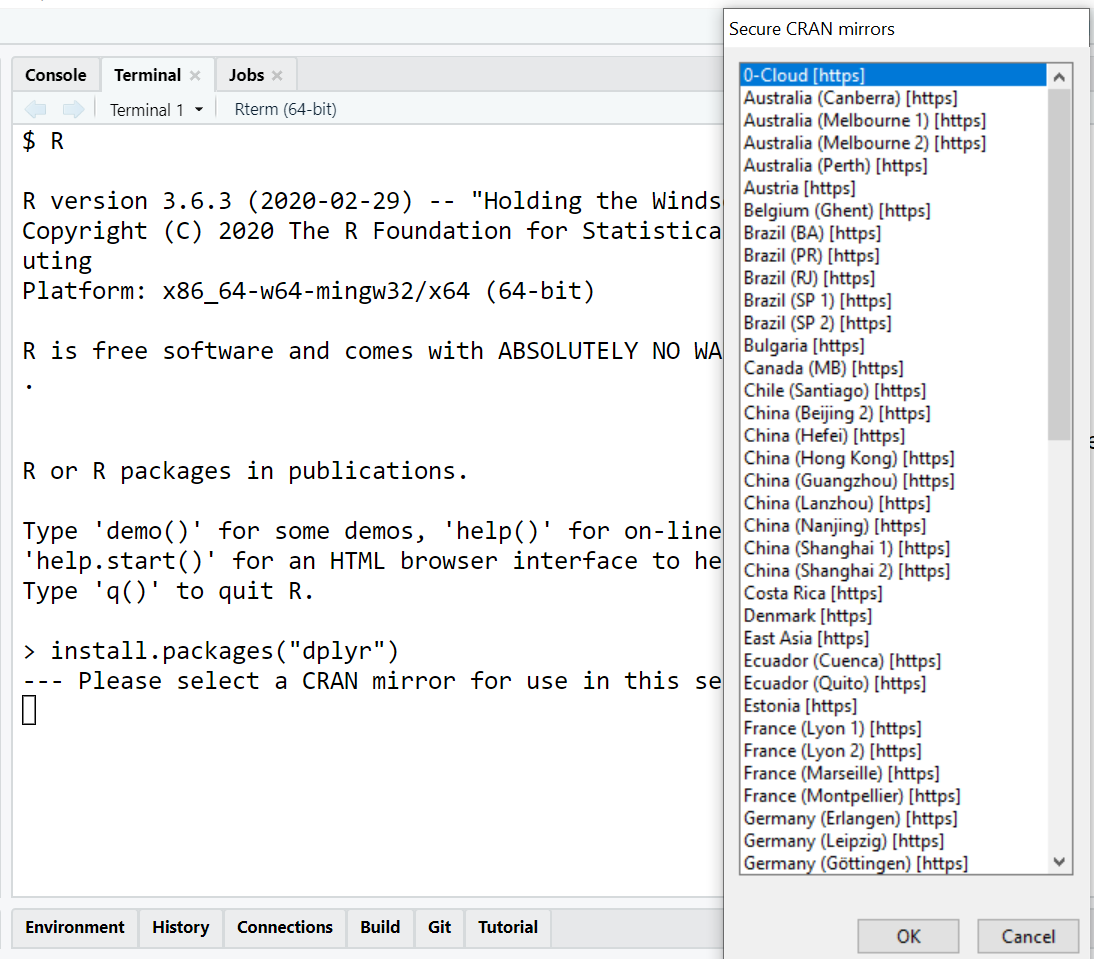
The most common way to install a package is by using the built-in install.packages() function. This function communicates with a CRAN mirror to download and install the package files to your local library.
To install a single package, such as the popular data manipulation tool dplyr, you would enter the following command in your console:
install.packages("dplyr")
It is crucial to remember that the package name must be enclosed in quotation marks. If you omit the quotes, R will look for an object named dplyr rather than searching the repository for a package of that name.
Installing Multiple Packages Simultaneously
Efficiency is key in software development. If you are setting up a new environment, you likely need more than one tool. R allows you to install multiple packages in a single line of code by passing a character vector to the function:
install.packages(c("ggplot2", "tidyr", "readr", "purrr"))
This command tells R to iterate through the list, checking for dependencies for each package and installing them in sequence. This is far more efficient than running individual commands for every library you require.
Utilizing the RStudio Graphical User Interface (GUI)
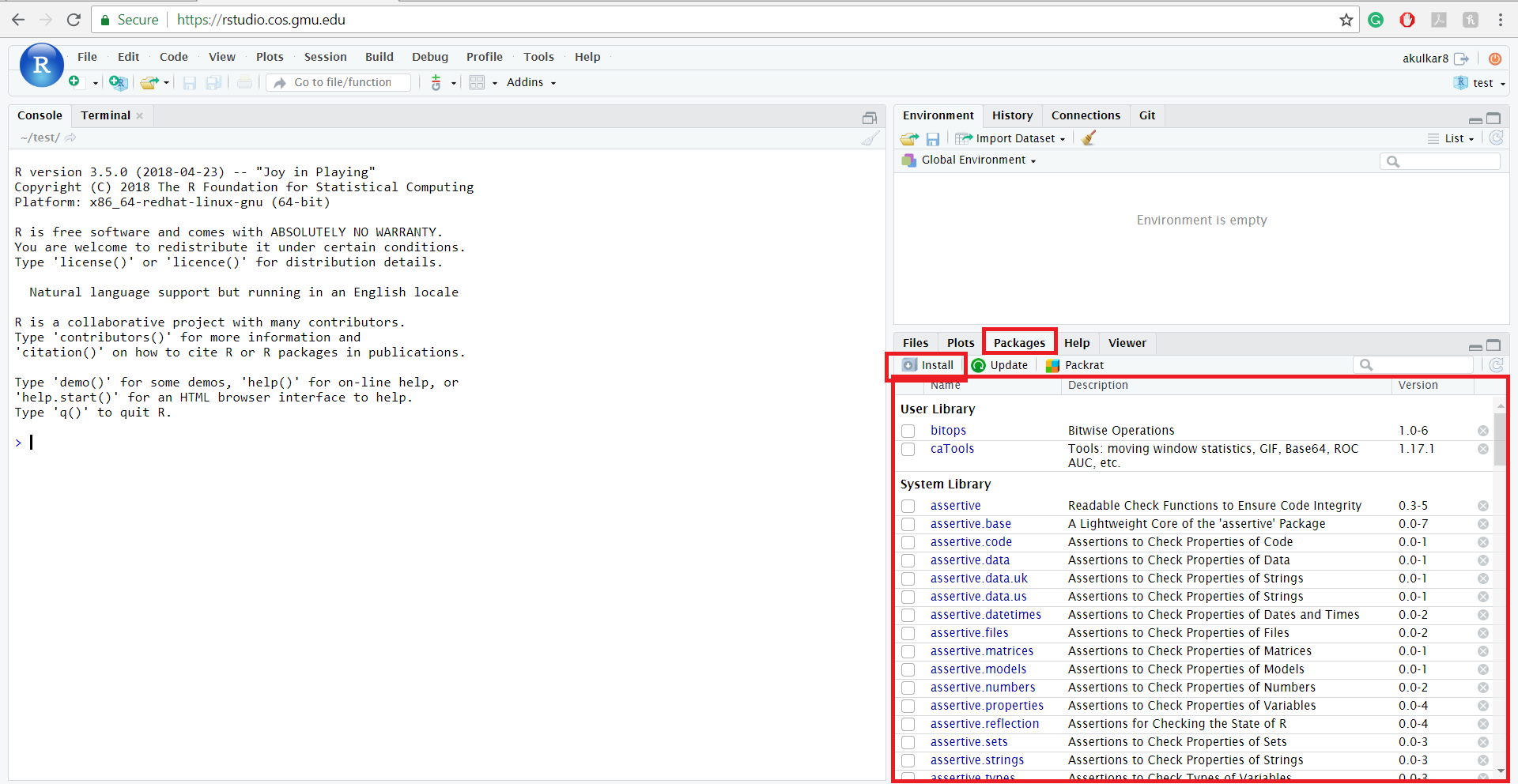
For those who prefer a visual approach, RStudio offers a user-friendly interface that abstracts the command line. In the “Files, Plots, Packages, Help” pane (usually in the bottom right), you will find a “Packages” tab.
Clicking the “Install” button opens a dialog box where you can type the names of the packages you want. RStudio also provides auto-complete suggestions, which is helpful if you are unsure of the exact spelling. This method still executes install.packages() in the background, but it provides a helpful layer of visual confirmation and allows you to choose your library path easily.
Advanced Installation Techniques
As your projects become more complex, you may find that the standard CRAN versions are insufficient. You might need a specific version of a package to maintain reproducibility or a development version to access a specific new feature.
Installing from GitHub via remotes and devtools
GitHub packages are not hosted as pre-compiled binaries like those on CRAN; they are hosted as source code. To install these, you need a tool that can bridge the gap. The devtools package (and its more lightweight counterpart, remotes) is the industry standard for this task.
First, you must install the helper package:
install.packages("remotes")
Then, you can install a package directly from a developer’s repository using the install_github function. The syntax follows a “developer/repository” format:
remotes::install_github("tidyverse/ggplot2")
This method is essential for tech professionals who contribute to open-source projects or those who need to test experimental features before they are officially released.
Installing Packages from Source Files
In some enterprise or high-security environments, R servers may not have direct access to the internet. In these cases, you cannot query CRAN or GitHub directly. Instead, you must manually download the package source file (usually a .tar.gz for Linux/macOS or a .zip for Windows) and install it locally.
The command for this is:
install.packages("C:/path/to/package/file.tar.gz", repos = NULL, type = "source")
By setting repos = NULL, you tell R not to look at an online repository, and type = "source" instructs it to compile the code from the provided file.
Managing Dependencies and Versioning
A common headache in software engineering is “dependency hell”—when Package A requires Package B version 1.0, but Package C requires Package B version 2.0. R handles dependencies automatically during the install.packages() process by default (using the argument dependencies = TRUE).
However, for professional-grade projects, relying on the latest version of every package can lead to broken code when packages update. Tools like renv (Reproducible Environments) allow developers to create a localized library for a specific project. This ensures that everyone working on the project uses the exact same versions of every package, drastically improving digital security and project stability.
Troubleshooting and Best Practices for Package Management
Installing software rarely goes perfectly every time. Understanding how to troubleshoot and maintain your R library is what separates a novice from a professional.
Common Installation Errors
One of the most frequent errors encountered, especially on Linux systems, is a “non-zero exit status.” This usually indicates that a system-level dependency is missing. For example, if you are installing a package that handles XML data, you might need to install the libxml2 library on your operating system before R can successfully compile the package.
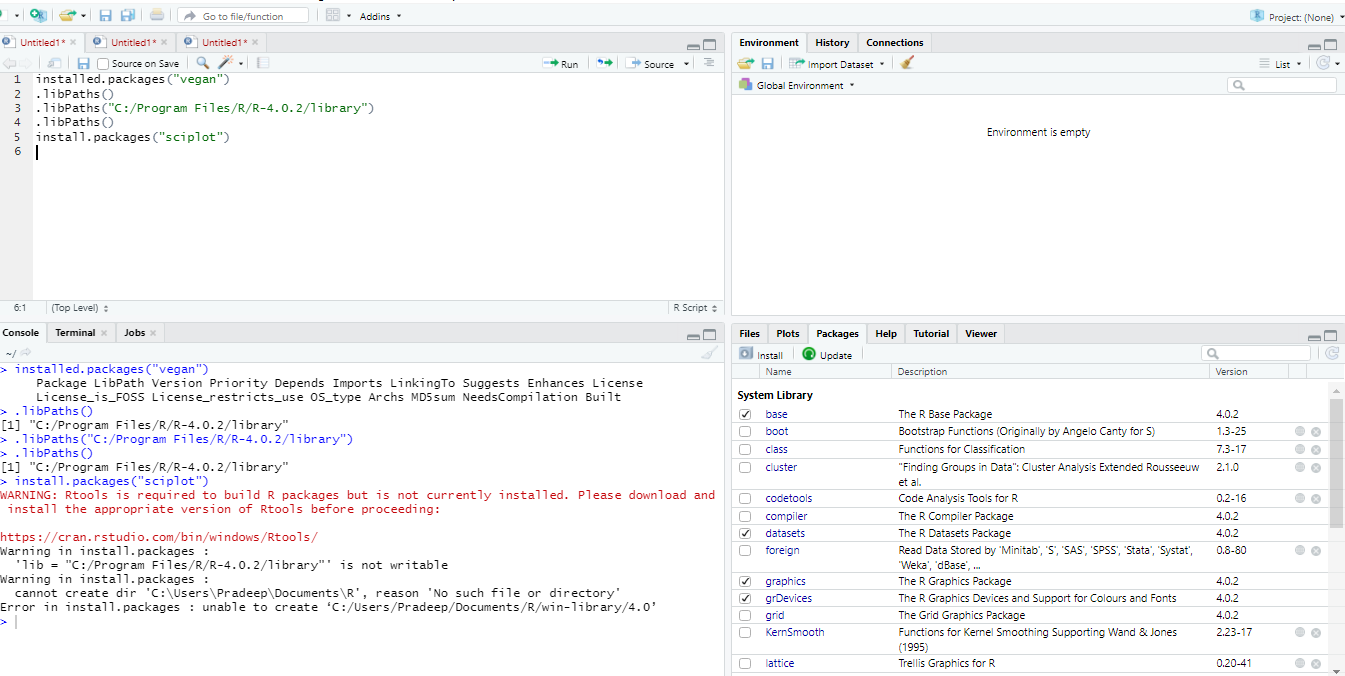
Another common issue is the “permission denied” error. This happens when R tries to write files to a folder that requires administrator privileges. A best practice is to ensure your R library is located in a folder where your user account has full read/write access.
Keeping Your Environment Updated
The tech world moves fast, and R packages are updated frequently to fix security vulnerabilities and improve performance. You can check for updates across your entire library using:
update.packages(ask = FALSE)
The ask = FALSE argument prevents R from prompting you for confirmation for every single package, streamlining the update process. It is advisable to run this periodically, though always be cautious when updating packages in the middle of a critical project.
Loading and Verifying Installation
Installing a package only puts the files on your hard drive. To actually use the functions within a script, you must load the package into your current R session using the library() function:
library(ggplot2)
If you want to check which packages are currently installed on your system and where they are located, you can use installed.packages(). For a cleaner look at your library paths, use .libPaths().
Conclusion: The Strategic Importance of Package Mastery
In the context of modern technology and data science, R packages are more than just scripts; they are the specialized tools that allow for complex problem-solving. Mastering the installation and management of these packages is a prerequisite for any serious data professional. From the stability of CRAN to the innovation found on GitHub, the ability to curate a precise technical environment is a hallmark of efficiency.
By understanding the underlying mechanisms of how R handles these external libraries—ranging from simple CLI commands to managing complex dependencies with renv—you ensure that your tech stack remains robust, reproducible, and secure. As AI and machine learning continue to integrate with R, staying adept at package management will remain a vital skill in the digital toolkit of any software or data professional.