In the intricate world of molecular biology, where the blueprint of life is encoded within DNA, a set of often-overlooked yet profoundly critical proteins stands as the unsung heroes of genetic organization: histone proteins. Far from mere structural components, histones are dynamic molecular architects, intricately coiling and packaging the vast stretches of DNA into the compact structure known as chromatin, which forms chromosomes. While their fundamental role lies in biology, the advent of sophisticated technologies has transformed our understanding of histones, propelling them to the forefront of biotechnology, drug discovery, and medical diagnostics. Understanding “what are histone proteins” today is less about a purely biological definition and more about appreciating their technological implications in shaping the future of medicine and synthetic biology.
These diminutive proteins, essential to all eukaryotic life, play a pivotal role not just in physically organizing DNA, but also in regulating gene expression, influencing everything from cellular differentiation to disease progression. Their study has become a vibrant intersection of biochemistry, genomics, computational science, and advanced imaging, firmly rooting their significance within the Tech category. From developing next-generation sequencing techniques to designing targeted therapies and leveraging artificial intelligence for epigenetic insights, the exploration of histone proteins is a testament to the power of technological innovation in unraveling biological complexity.
The Fundamental Role of Histones: DNA Packaging and Gene Regulation
At the core of cellular function, histone proteins perform a dual role: compacting the immense length of DNA to fit within the microscopic confines of a cell’s nucleus and regulating the accessibility of genes for transcription. This foundational understanding is critical for developing technologies that can manipulate or monitor these processes.
The Nucleosome: The Basic Unit of Chromatin Architecture
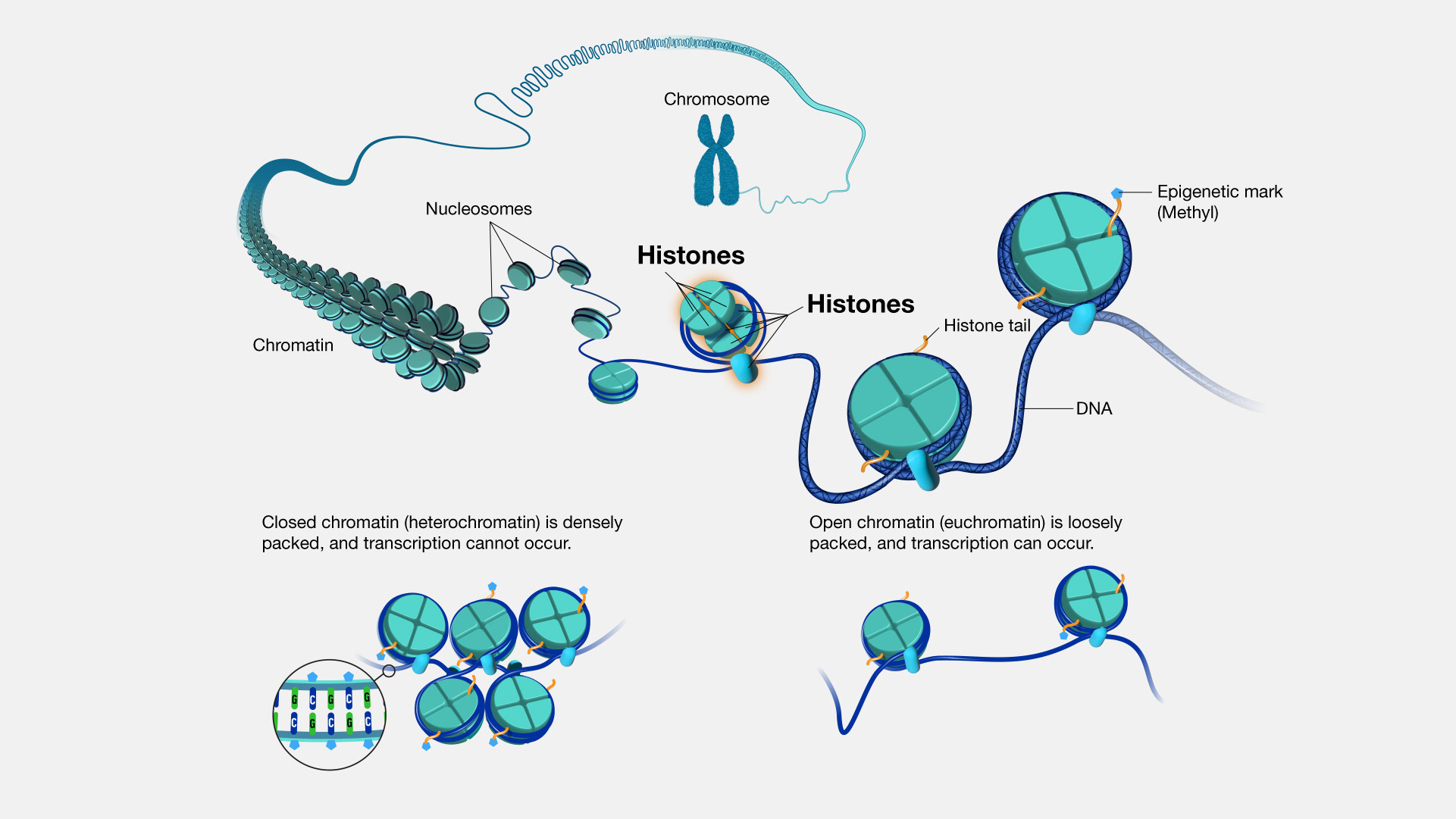
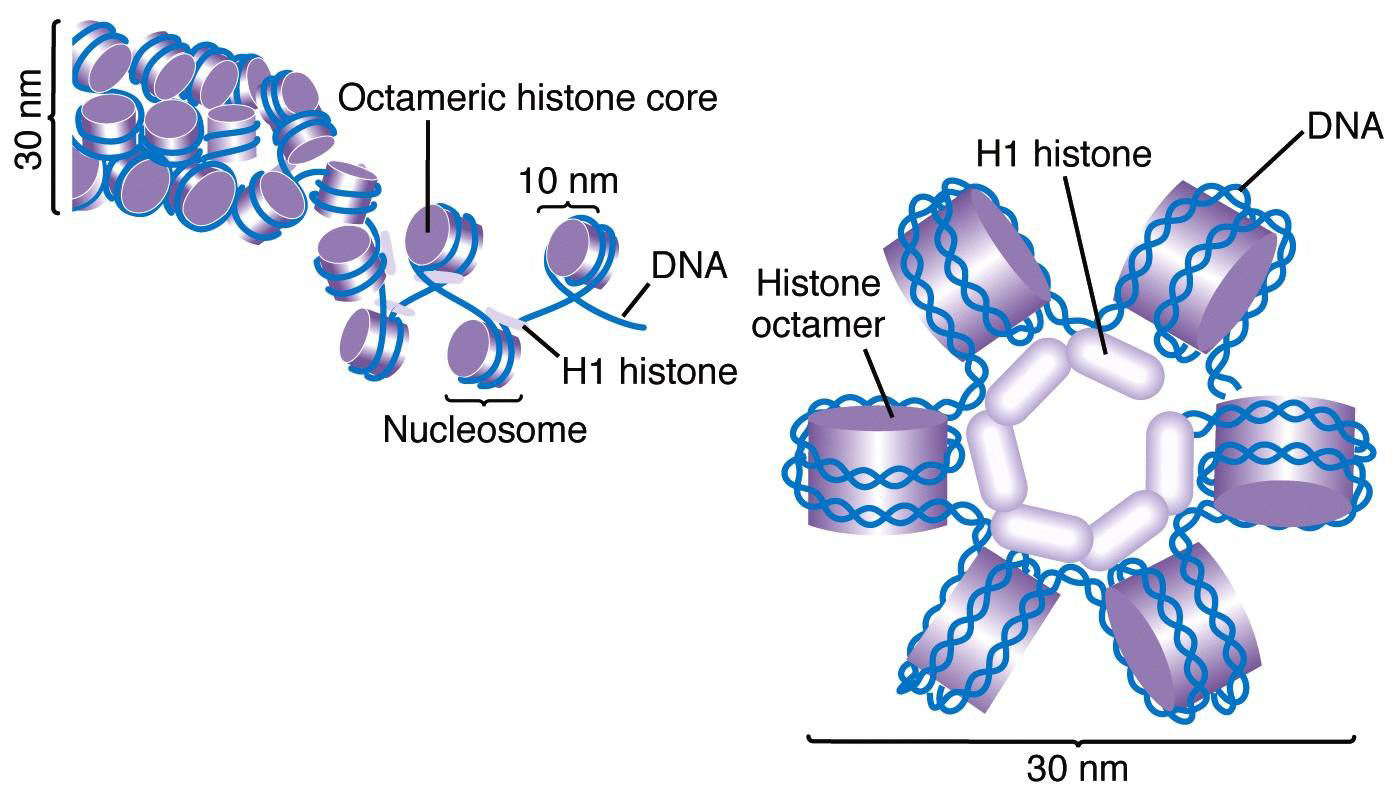
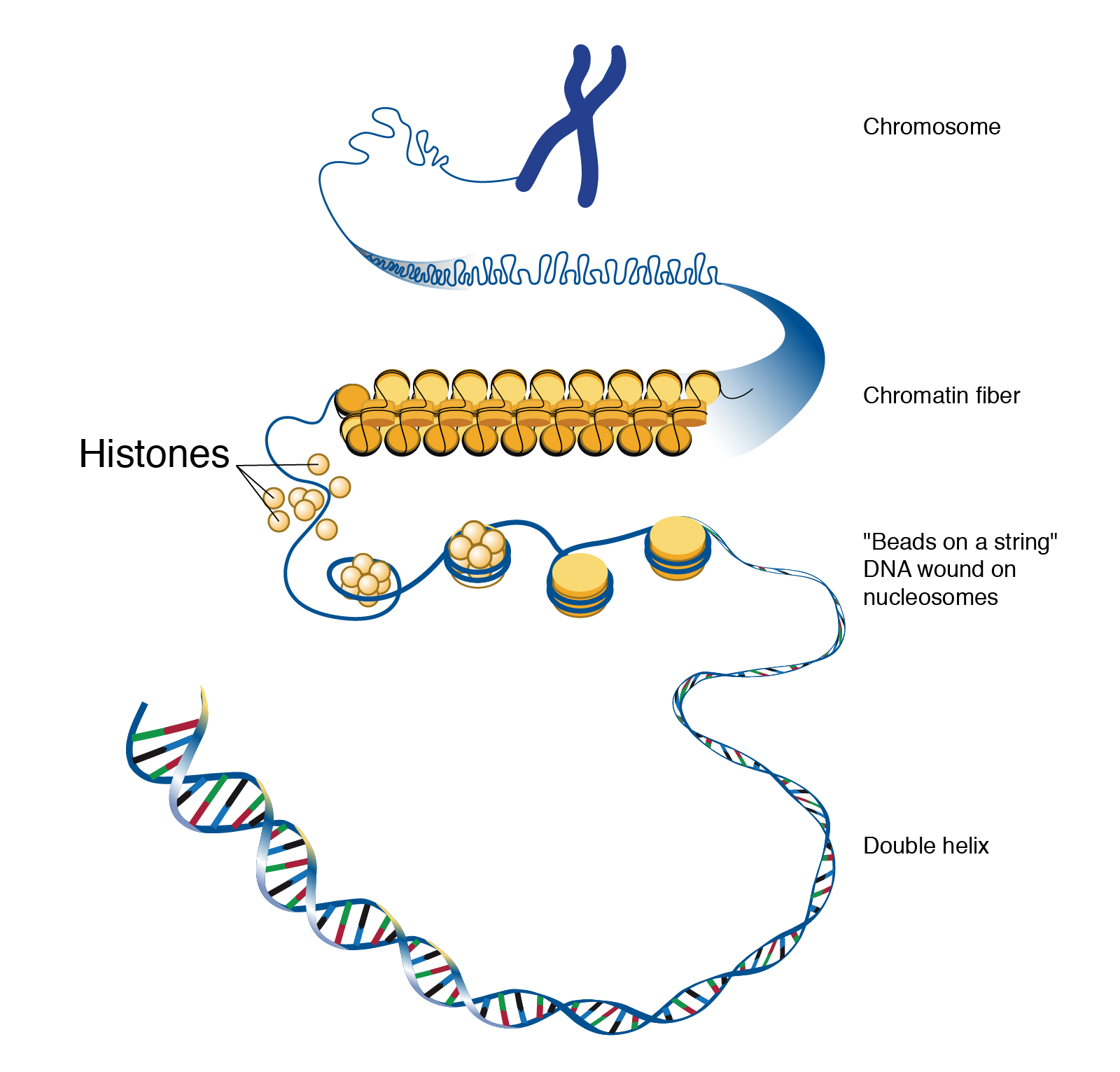
The human genome, if stretched out, would measure approximately two meters in length. To fit this colossal molecule into a nucleus just a few micrometers in diameter, DNA must be exquisitely organized. This organization begins with histone proteins. There are five main classes of histones: H1, H2A, H2B, H3, and H4. Histones H2A, H2B, H3, and H4 are known as the core histones. Two molecules of each of these four core histones assemble to form an octamer, a protein complex around which approximately 147 base pairs of DNA are wrapped, much like thread around a spool. This fundamental unit of DNA packaging is called a nucleosome. Nucleosomes are then further compacted into higher-order chromatin structures, a process often facilitated by histone H1, which acts as a linker histone.
The technological significance here lies in our ability to observe and model these structures. Advanced cryo-electron microscopy (Cryo-EM) and X-ray crystallography are cutting-edge technologies that have allowed scientists to visualize the atomic-level structure of nucleosomes and chromatin fibers, providing crucial insights into their assembly and dynamics. These structural revelations are indispensable for rational drug design and for engineering synthetic biological systems that mimic or manipulate natural chromatin. Computational tools further extend this capability, enabling simulations of chromatin dynamics under various conditions, which is paramount for predicting gene expression patterns.
Epigenetics: The Software of Gene Expression and Its Technological Implications
Beyond their structural role, histones are central to the field of epigenetics—the study of heritable changes in gene expression that occur without altering the underlying DNA sequence. Histones are subject to a myriad of post-translational modifications (PTMs), such as acetylation, methylation, phosphorylation, and ubiquitination, primarily on their amino-terminal tails that protrude from the nucleosome. These “histone codes” act as molecular switches, influencing whether specific regions of DNA are tightly packed and inaccessible (gene silencing) or loosely packed and accessible (gene activation).
The concept of the histone code is akin to the “software” layer that dictates how the “hardware” (DNA sequence) is read and utilized. Understanding and deciphering this epigenetic software is a major frontier in biotechnology. The ability to read, write, and erase these histone modifications offers unprecedented opportunities for therapeutic intervention. Technologies like ChIP-seq (Chromatin Immunoprecipitation sequencing), ATAC-seq (Assay for Transposase-Accessible Chromatin using sequencing), and Hi-C (High-throughput Chromosome Conformation Capture) are specialized genomic sequencing techniques that allow researchers to map these modifications and chromatin accessibility across entire genomes. These high-throughput technologies generate vast datasets that require sophisticated bioinformatics pipelines, machine learning algorithms, and AI tools for analysis and interpretation, firmly placing epigenetic research within the realm of Big Data and AI-driven biotechnology.
Technological Approaches to Unraveling Histone Mysteries
The study of histone proteins and their complex modifications has been revolutionized by a suite of advanced technological platforms. These tools enable scientists to probe chromatin structure and function with unprecedented resolution and scale, driving discoveries in basic science and translational medicine.
Advanced Sequencing and Epigenomics Platforms
The era of genomics has given rise to sophisticated tools for dissecting the epigenome. ChIP-seq, for instance, allows researchers to identify regions of DNA associated with specific histone modifications or proteins, providing a genome-wide map of epigenetic marks. ATAC-seq offers insights into chromatin accessibility, revealing regions where DNA is open for gene transcription. These methods leverage next-generation sequencing technologies, which have seen exponential increases in throughput and reductions in cost, making large-scale epigenomic studies feasible. The data generated from these platforms is massive and requires robust computational infrastructure for storage, processing, and analysis. This involves advanced algorithms for peak calling, differential analysis, and data visualization, often integrated into user-friendly software packages that form part of modern bioinformatics toolkits.
Computational Biology and AI in Histone Research
The sheer volume and complexity of epigenomic data necessitate advanced computational approaches, including artificial intelligence and machine learning. AI algorithms are increasingly being used to predict histone modification patterns, identify regulatory elements, and even model the three-dimensional structure of chromatin based on epigenetic cues. For example, deep learning models can be trained on ChIP-seq data to learn intricate relationships between histone marks and gene expression, enabling more accurate predictions of gene activity. Furthermore, computational simulations can model the dynamic interactions between DNA, histones, and regulatory proteins, helping to understand how changes in histone modifications propagate through the chromatin landscape. These AI-driven tools are not just analytical; they are becoming predictive and even generative, accelerating hypothesis generation and experimental design in histone research.
High-Resolution Imaging Techniques
While sequencing provides linear information about the epigenome, high-resolution imaging technologies offer spatial insights into chromatin organization within the nucleus. Techniques like super-resolution microscopy (e.g., STORM, PALM) can visualize individual nucleosomes and histone modifications in live cells with nanoscale precision, providing a dynamic view of chromatin structure. Cryo-electron microscopy (Cryo-EM) has been instrumental in resolving the atomic structures of nucleosomes and larger chromatin assemblies, offering blueprints for molecular interactions. These imaging technologies provide a critical bridge between biochemical analyses and cellular function, allowing researchers to observe the effects of histone modifications on chromatin condensation and gene accessibility in real time. The development of advanced image processing software and machine vision algorithms is crucial for extracting meaningful biological information from these complex datasets.
Histones at the Forefront of Biomedical Innovation
The deepened understanding of histone proteins and their epigenetic modifications, powered by technological advancements, has profound implications for various sectors of biomedical innovation, particularly in drug discovery, gene editing, and diagnostic development.
Drug Discovery and Therapeutic Targets
The reversible nature of histone modifications makes them attractive targets for therapeutic intervention. Companies are developing small molecule inhibitors that target enzymes responsible for adding or removing histone marks. For example, histone deacetylase (HDAC) inhibitors are a class of drugs already approved for certain cancers, working by increasing histone acetylation, which generally promotes gene expression and can induce cell differentiation or apoptosis in cancer cells. Similarly, inhibitors targeting histone methyltransferases (HMTs) or demethylases (HDMs) are in various stages of clinical development. The process of identifying, validating, and optimizing these epigenetic drugs relies heavily on high-throughput screening technologies, computational chemistry, and advanced cellular assays. Furthermore, personalized medicine approaches aim to tailor these therapies based on an individual’s specific epigenetic profile, as revealed by advanced diagnostic technologies.
Gene Editing and Synthetic Biology
The advent of powerful gene editing technologies like CRISPR-Cas systems has opened new avenues for manipulating the genome, and histones are increasingly recognized as key players in this space. While CRISPR primarily targets DNA sequences, its efficacy and specificity can be influenced by chromatin accessibility and histone modifications. Researchers are now exploring ways to combine gene editing with epigenetic engineering, for example, by delivering CRISPR-based tools that can specifically alter histone modifications at target loci, thereby enhancing or repressing gene expression without altering the DNA sequence itself. This emerging field of “epigenome editing” holds immense promise for treating genetic diseases where gene expression levels, rather than just DNA sequence, are perturbed. Synthetic biology also seeks to engineer novel biological systems, and designing precise control over gene expression often involves mimicking or re-engineering histone-DNA interactions.
Diagnostic Biomarkers for Disease
Abnormal histone modification patterns are increasingly recognized as hallmarks of various diseases, including cancer, neurodegenerative disorders, and metabolic diseases. This makes histone modifications powerful candidates for diagnostic and prognostic biomarkers. For instance, specific patterns of histone acetylation or methylation in circulating tumor DNA (ctDNA) or blood cells could serve as early detection markers for cancer or indicators of treatment response. The development of highly sensitive and specific assays, often leveraging advanced microfluidics and molecular barcoding technologies, is crucial for translating these discoveries into clinical diagnostics. The integration of AI and machine learning can help in analyzing complex epigenetic profiles to identify robust biomarkers, moving towards precision diagnostics that inform tailored treatment strategies.
Challenges and Future Frontiers in Histone-Related Technologies
Despite the immense progress, several challenges remain in fully harnessing the potential of histone proteins in technology and medicine. Addressing these challenges will define the future frontiers of research and development.
Complexity of the Epigenome: Data Integration and Interpretation
The sheer number of possible histone modifications, their combinatorial nature, and their dynamic interplay with other epigenetic mechanisms (like DNA methylation and non-coding RNAs) create an incredibly complex regulatory landscape. Integrating disparate epigenomic datasets—from ChIP-seq and ATAC-seq to RNA-seq and proteomic data—to build comprehensive models of gene regulation is a major computational and analytical challenge. Future technologies will need to focus on multi-omics data integration platforms and advanced AI algorithms capable of discerning meaningful biological patterns from this vast and noisy data.
Ethical Considerations in Epigenetic Engineering
As our ability to manipulate histone modifications and the epigenome grows, so do the ethical considerations. Epigenetic changes can be heritable, raising questions about germline editing and its long-term consequences. The potential for unintended side effects from broad-spectrum epigenetic drugs or targeted epigenome editing also requires careful scrutiny. Developing robust regulatory frameworks and engaging in public discourse are essential to ensure responsible technological advancement in this sensitive area.
Personalized Medicine and the Epigenetic Revolution
The ultimate goal for many histone-related technologies is to enable truly personalized medicine. By profiling an individual’s unique epigenetic landscape, clinicians could potentially predict disease risk, select the most effective therapies, and monitor treatment responses with unprecedented precision. The future will likely see the development of portable, high-throughput epigenetic profiling devices and AI-powered diagnostic platforms that can quickly analyze a patient’s epigenome and recommend tailored interventions. This “epigenetic revolution” in personalized medicine hinges on continued innovation in both the fundamental understanding of histones and the development of accessible, reliable technological solutions.
In conclusion, histone proteins, though small, are mighty. Their fundamental role in DNA packaging and gene regulation makes them central to life itself. However, it is through the lens of modern technology—from advanced sequencing and imaging to computational biology and AI—that their true potential in revolutionizing medicine, diagnostics, and synthetic biology is being realized. As we continue to push the boundaries of what is technologically possible, our understanding and manipulation of these molecular architects will undoubtedly unlock new frontiers in human health and beyond, firmly establishing “what are histone proteins” as a question with ever-expanding technological answers.
aViewFromTheCave is a participant in the Amazon Services LLC Associates Program, an affiliate advertising program designed to provide a means for sites to earn advertising fees by advertising and linking to Amazon.com. Amazon, the Amazon logo, AmazonSupply, and the AmazonSupply logo are trademarks of Amazon.com, Inc. or its affiliates. As an Amazon Associate we earn affiliate commissions from qualifying purchases.